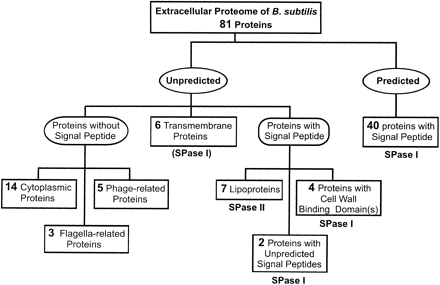
Overview of identified and predicted extracellular proteins of B. subtilis 168. Extracellular proteins, as listed in Table 1, were classified as predicted or unpredicted with respect to their extracellular localization on the basis of previous genome-based predictions of signal peptides and retention signals (Tjalsma et al. 2000). Proteins with signal peptides are further classified on the basis of their cleavage sites for SPase I or II, and the presence of cell-wall-binding domains. Note that potential pre-proteins with an SPase I cleavage site and predicted transmembrane segments were excluded from previous predictions. XynD, which has a predicted SPase II cleavage site in addition to a potential transmembrane segment, is counted as a transmembrane protein; both predictions are, however, probably wrong (see Results section and Table 1A). The indicated numbers correspond to numbers printed in italics in the Results section.











