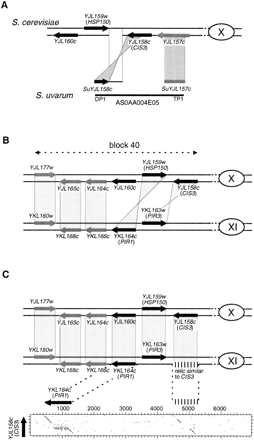
Local gene inversions. Symbols are the same as in Figure 1. (black arrows) Genes that are members of the same gene family. (A) Local gene inversion of SuYJL158c. The chromosomal neighborhood of YJL158c (CIS3) in S. cerevisiaechromosome X is presented at the top of the figure. At the bottom, hits are depicted that were obtained from bothBLASTX (dark shading) and BLASTN (light shading) comparisons of the two S. uvarum paired RSTs (AS0AA004E05DP1 and AS0AA004E05TP1) with the S. cerevisiaegenome. (B) Structure of the duplicated block 40 in S. cerevisiae. Only those genes being duplicated between and/or within the two blocks are represented. The shaded areas between genes represent the BLASTP-based identification of the paralogous partners (Wolfe and Schields 1997). (C) Identification of a relic of YJL158c. The relic on S. cerevisiae chromosome XI is symbolized by the small vertical lines and the corresponding DNA dot matrix is presented below (stringency/window set a 15/23). The shaded area corresponds to the new paring of duplicates in block 40.











