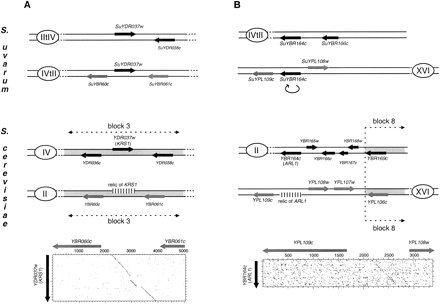
Loss of duplicated genes within S. cerevisiae ancestral duplication blocks. Symbols are the same as in Fig. 1. Shaded parts of the chromosomes correspond to duplication blocks in S. cerevisiae (Wolfe and Schields 1997). The relics ofYDR037w (KRS1) and YBR164c (ARL1) identified in theS. cerevisiae genome are symbolized by the small vertical lines. (A) Loss of the second copy of YDR037w within the S. cerevisiae duplication block 3 on chromosome II:YDR037w is a singleton in the S. cerevisiae genome whereas it is duplicated in S. uvarum. One of the two copies was isolated next to YDR038c, as expected, whereas the second copy was identified between SuYBR060c and SuYBR061c. A relic of the second copy of YDR037w was detected by DNA dot matrix by aligning the sequence of the intergenic region betweenYBR060c and YBR061c with the sequence ofYDR037w. The stringency/window parameters of the dot matrix were set at 15/23. (B) Loss of the second copy ofYBR164c on the border of the S. cerevisiaeduplication block 8 on chromosome XVI. YBR164c is a singleton in S. cerevisiae whereas it is duplicated in S. uvarum. One of the two copies was isolated associated toSuYBR166c whereas the other copy is located betweenSuYPL108w and SuYPL109w. In S. cerevisiae SuYPL108w and SuYPL109w lie outside of the duplicated block 8. In S. uvarum, the second copy ofSuYBR164c is inverted with respect to the centromere (as symbolized by the curved arrow), as is the case for the whole of block 8 in S. cerevisiae. A relic of the second copy ofYBR164c was identified in the S. cerevisiae genome in the intergenic region between SuYPL108w andSuYPL109w. The stringency/window parameters of the dot matrix were set at 11/19.











