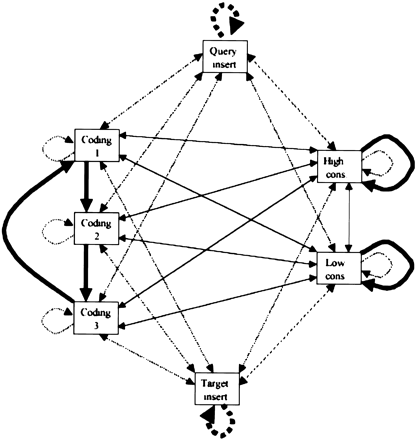
Finite state machine depiction of seven state aligner. This is a pairwise HMM. Transitions between states are associated either with aligning a pair of nucleotides (solid arrows) or aligning a single nucleotide with a gap (broken line arrows). The most likely transition out of each state is shown by a thick arrow. Thin arrows represent possible, but unlikely, transitions. In coding regions, the most likely path will typically cycle between coding states 1, 2, and 3. In strongly conserved regions, the most likely path will typically stay in the high conservation state. In weakly conserved regions, the most likely path will typically stay in the low conservation state. Short inserts follow the broken line arrows in the coding and conserved states. Long inserts transition into the query or target insert states.











