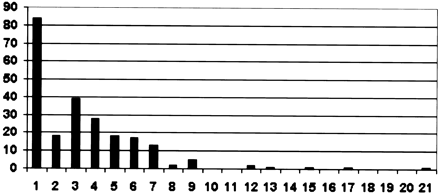
Synteny summary graph. The horizontal axis is how many pieces a C. briggsae clone needed to be broken into for maximal alignment with the C. elegans genome. The vertical axis is the number of clones broken into that many pieces. There were 229 clones, and the average clone size was 34,722 bases. Clones were broken up if the best aligning region of one 2000-base region of the C. briggsaeclone aligned to a spot 50,000 bases or more away or on a different chromosome from where the previous 2000-base region of the C. briggsae clone aligned best. The 2000 base regions overlapped by 1000 bases. The average length of a broken-up region was 8553 bases. The bimodal distribution suggests that ∼40% of the genome is resistant to rearrangements and the rest is not.











