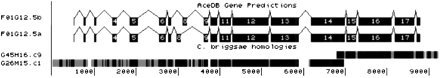
Tracks display of let-2, a gene that is alternatively spliced in a developmentally regulated manner. The C. briggsaesequence is split between two overlapping cosmids. Note the relatively high level of conservation in the introns compared withunc-47. (The exons numbered 8 on the top and bottom splicing diagrams are numbered 9 and 10 in the text and in Table 5. This difference is due to Tracks display not including exons in the UTR regions and numbering the exons in each isoform separately.)











