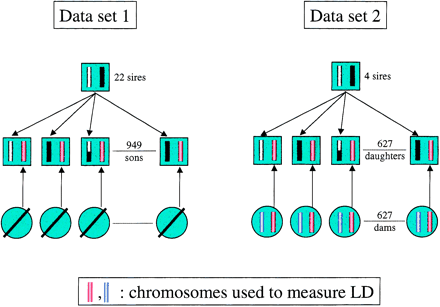
(Left) Data set 1: Data set 1 corresponds to a GDD described previously, that is, a series (22) of paternal half-brother families with their sires for a total of 22 + 949 bulls (Coppieters et al. 1998). The dams of data set 1 are not genotyped. The marker linkage phase of the founder sires are determined from the genotypes of their respective sons as described (Georges et al. 1995). Assuming known marker phase of the sire, the most likely genotypes of paternal (black, white, or recombinant) and maternal (red) gametes transmitted to the son can be inferred. The genotypes of the maternal gametes (red) were used to measure LD. (Right) Data set 2: Data set 2 corresponds to a daughter design, that is, a series (4) of paternal half-sister families with their sires for a total of 624 daughters. The 624 dams of data set 2 were genotyped as well. The marker-linkage phase of the founder sires are determined from the genotypes of their respective daughters as described, exploiting the available genotype information from the dams (Georges et al. 1995). Assuming known marker phase of the sire, the most likely genotypes of paternal (black, white, or recombinant) and maternal (red) gametes transmitted to the daughters can be inferred. The genotypes of the maternal gametes (red) as well as their complement (blue) were used to measure LD.











