Zebrafish Comparative Genomics and the Origins of Vertebrate Chromosomes
Abstract
To help understand mechanisms of vertebrate genome evolution, we have compared zebrafish and tetrapod gene maps. It has been suggested that translocations are fixed more frequently than inversions in mammals. Gene maps showed that blocks of conserved syntenies between zebrafish and humans were large, but gene orders were frequently inverted and transposed. This shows that intrachromosomal rearrangements have been fixed more frequently than translocations. Duplicated chromosome segments suggest that a genome duplication occurred in ray-fin phylogeny, and comparative studies suggest that this event happened deep in the ancestry of teleost fish. Consideration of duplicate chromosome segments shows that at least 20% of duplicated gene pairs may be retained from this event. Despite genome duplication, zebrafish and humans have about the same number of chromosomes, and zebrafish chromosomes are mosaically orthologous to several human chromosomes. Is this because of an excess of chromosome fissions in the human lineage or an excess of chromosome fusions in the zebrafish lineage? Comparative analysis suggests that an excess of chromosome fissions in the tetrapod lineage may account for chromosome numbers and provides histories for several human chromosomes.
As the first human chromosomes become completely sequenced (Dunham et al. 1999; Hattori et al. 2000), attention turns to a genome-wide understanding of sequence function and the mechanisms of genome evolution. The study of human genome evolution profits from comparative analysis. Progress relating human gene maps to those of other mammals has been rapid (DeBry and Seldin 1996; Schibler et al. 1998; O'Brien and Stanyon 1999; Watanabe et al. 1999; Murphy et al. 2000), but investigation of outgroups to mammals is necessary to distinguish shared, derived features of mammalian genomes from features inherited from ancestral genomes. A useful outgroup is the chicken (Groenen et al. 2000), whose lineage separated from that of mammals ∼310 million years ago (mya; Kumar and Hedges 1998). Chicken, however, has a complex genome consisting of 39 chromosome pairs, nearly twice the number of mouse (20 pairs), and many of these are microchromosomes (Bloom et al. 1993). Another valuable outgroup is the zebrafish Danio rerio. Thousands of mutations that disrupt embryonic development have been discovered in zebrafish, and their analysis can reveal conserved gene functions and provide models for human disease and congenital malformation (Brownlie et al. 1998; Wang et al. 1998; Donovan et al. 2000; Feldman et al. 2000). In addition, comparative analysis of zebrafish and mammalian gene maps can help clarify the origin of the human genome.
Prior analysis revealed several chromosome regions (Amores et al. 1998;Postlethwait et al. 1998; Gates et al. 1999) that were syntenic in the last common ancestor of human and zebrafish ∼450 mya (Kumar and Hedges 1998). Such conserved syntenies have expedited the molecular identification of zebrafish mutations (Karlstrom et al. 1999; Donovan et al. 2000; Miller et al. 2000). Earlier analyses, however, did not address the extent to which gene orders are preserved within blocks of conserved synteny. It has been observed that rearrangements between chromosomes have occurred about four times more frequently than inversions within a chromosome, at least in mammals (Ehrlich et al. 1997). If this mechanism of chromosome evolution holds for vertebrates in general, then conserved syntenies will be relatively short, but the orders of genes within conserved syntenies should be mostly preserved. Alternatively, if inversions are more frequently fixed in vertebrate evolution than are translocations, then blocks of conserved syntenies will be large, but the orders of loci within conserved blocks will be rearranged. With the increased resolution provided by human radiation hybrid maps (Stewart et al. 1997), we can now test these two hypotheses by comparing corresponding regions of the zebrafish and human genomes.
Our earlier comparative analysis revealed an apparently genome-wide duplication in the ancestry of zebrafish (Amores et al. 1998;Postlethwait et al. 1998; Gates et al. 1999). Here we address the ways in which duplicated chromosome segments have evolved. A key feature of the evolution of duplicated chromosomes is the fraction of genes retained as duplicate pairs. To estimate this fraction for zebrafish, we have compared the content of several duplicated chromosome segments.
Comparative genomics among teleost fish suggests that the genome-wide duplication detected in zebrafish probably occurred before the divergence of zebrafish, pufferfish, and medaka lineages (Amores et al. 1998; Meyer and Malaga-Trillo 1999; Meyer and Schartl 1999; Aparicio 2000; Naruse et al. 2000) at the base of the teleost radiation, >100 mya (Santini and Tyler 1999). Despite this duplication, zebrafish has about the same number of chromosomes as humans (25 vs. 23 pairs), rather than twice as many, as would be expected in the absence of chromosome rearrangements. At least two hypotheses might account for this situation. First, there could have been an excess of chromosome fusions in the fish lineage compared to the human lineage. Alternatively, there could have been an excess of chromosome fission events in the human lineage because it diverged from the lineage of ray fin fishes. We have examined several specific zebrafish chromosomes and their mammalian orthologs to address these two hypotheses.
To help understand the evolution of mammalian genomes, we have placed a group of 54 cloned genes and expressed sequence tags (ESTs) on a previously established meiotic mapping panel (Kelly et al. 2000; Woods et al. 2000). Our results indicate the content of certain chromosomes in the genome of the last common ancestor of teleosts and mammals and provide evidence that there has been an excess of chromosome fissions over chromosome fusions in the mammalian lineage.
RESULTS
Evolution of Human Chromosome 17 and its Zebrafish Duplicates
To illuminate the evolution of human chromosome 17 (Hsa17), we searched the zebrafish databases for ESTs and cloned genes for loci apparently orthologous to loci on Hsa17 (as determined by reciprocal BLAST comparisons) and then mapped these zebrafish loci on the HS meiotic mapping panel (Table 1, available as supplementary material athttp://www.genome.org; Kelly et al. 2000; Woods et al. 2000). The results showed that most orthologs of Hsa17 map to LG3, LG5, LG12, and LG15.
Linkage Group 3
Among the 68 coding loci mapped to LG3 in the HS panel in previous work (Kelly et al. 2000; Woods et al. 2000) and the 19 added in the current work, 29 had putative orthologs on Hsa17 (Fig.1A). Adding two other loci mapped on the MOP panel (Postlethwait et al. 1998) gives 31 putative orthologs of Hsa17 genes on LG3. These loci were distributed along the length of Hsa17 roughly according to its gene density (http://www.ncbi.nlm.nih.gov/genemap/), except for the region between ∼150–300 cR3000. We conclude that a portion of LG3 is orthologous to most of human chromosome 17. Hsa17 loci are all (or nearly all) found on a single chromosome in cat (Murphy et al. 2000), rat (Watanabe 1999), and mouse (DeBry and Seldin 1996). We conclude that loci on most of the chromosome segment that is now Hsa17 have remained syntenic in the lineage leading to mammals since the divergence of human and zebrafish genomes 450 mya.
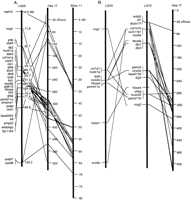
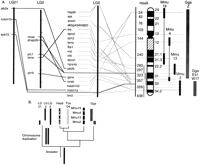
Zebrafish chromosome segments orthologous to Hsa17. (A) LG3 has many loci orthologous to mammalian genes distributed along Hsa17 and mouse chromosome Mmu11. Despite the large block of conserved synteny, multiple intrachromosomal rearrangements have altered gene orders in fish and mammalian chromosomes. (B) LG12 also contains many loci orthologous to Hsa17 genes, and many of these have duplicate copies on LG3. Accession nos. are listed in Supplementary Data at http://www.genome.org. See Woods et al. (2000) for a complete map for markers in this and following figures.
This extensive and very ancient block of conserved syntenic loci on LG3 provides a test of the prediction that translocations should be much more common than inversions in vertebrate evolution (Ehrlich et al. 1997). If inversions are infrequent with respect to translocations, then the orthologous zebrafish and human chromosomes should be colinear. Figure 1A compares gene orders along the corresponding chromosome segments of human and zebrafish. The results show that loci located in the same bin in the zebrafish HS mapping panel can reside at different locations in the human map. This shows that, while translocations have not disrupted the block of genes, multiple inversions have been fixed in populations. For comparative purposes, Figure 1A relates Hsa17 and mouse chromosome 11 (Mmu11) for loci orthologous to those in our data set. The comparison reveals several inversions of gene orders between the two mammals, and in total, there are six conserved segments ordered (CSO; Murphy et al. 2000) along Hsa17 in which the orders of loci are conserved in mouse and human. Not surprisingly, more internal chromosome rearrangements have been fixed in the 450 million years since zebrafish and human genomes diverged than in the ∼100 million years separating the mouse and human genomes (Kumar and Hedges 1998;Bromham et al. 1999; but see Bromham et al. 2000).
Linkage Group 12
A large block of zebrafish loci that are putatively orthologous to Hsa17 genes is also found on LG12 (Fig. 1B). Are LG3 and LG12 derived from the fragmentation of an Hsa17 ortholog in the ray-fin fish lineage? Because the human orthologs of LG12 genes are distributed along the human chromosome approximately like those of LG3, a simple translocation seems unlikely. Another possible explanation is that LG3 and LG12 are duplicates of each other, both being orthologous to Hsa17. This supposition is supported by the finding that zebrafish has two orthologs of several single-copy genes on Hsa17, and in most of these cases, one duplicate copy maps to LG3 and the other to LG12. These cases include duplicates of the human genes SOX9,COL1A1, HBOA, DLX7, RARA,HOXB1, HOXB5, HOXB6, HOXB8,PSMD11, and NOG. In addition, these two zebrafish chromosomes have duplicate loci for HBA from Hsa16 andHBB from Hsa11. (In mouse, as in zebrafish, Hba is syntenic with orthologs of the 11 duplicated loci mentioned above.) These data are consistent with the hypothesis that portions of LG3 and LG12 are derived from a chromosome duplication event.
The results discussed above showed that numerous intrachromosomal rearrangements have rearranged LG3 with respect to its human ortholog Hsa17. Have any of these rearrangements occurred in the zebrafish lineage since the LG3/LG12 chromosome duplication event? Figure 1B shows that the relative order of duplicate loci on LG3 and LG12 are not colinear, indicating that a number of inversions have occurred in the >100 million years since the duplication event. The density of duplicate markers is insufficient to make a quantitative estimate of relative rates in different lineages.
Linkage Groups 5 and 15
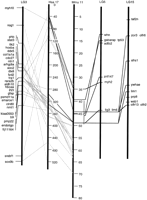
At least eight putative orthologs of Hsa17 loci are on LG5 and at least 10 are on LG15 (Fig. 2). These loci occupy a restricted portion of Mmu11 between 35 and 49 cM, whereas the putative Hsa17 orthologs on LG3 and LG12 occupy the two flanking regions on Mmu11. The location of lim1 and lim6(duplicated orthologs of human LHX1 [Hsa17_302.47 cR3000] and mouse Lhx1 [Mmu11_48.0 cM]) on LG15 and LG5, respectively, further supports the idea that LG5 and LG15 contain duplicate chromosome segments. Two positions on LG15 contain clusters of zebrafish olfactory receptor genes (Weth et al. 1996; Barth et al. 1997), and there is a large cluster of closely related olfactory receptors on Hsa17p13.3. Orthologies, however, are difficult to determine within this very large and divergent gene family. Hsa17p13.1 contains a cluster of seven or more myosin heavy chain genes, and we mapped putative orthologs of these to LG3 and LG5. If gene clusters like these existed in the last common ancestor of zebrafish and human, then it is possible, or even likely, that there would be no true orthologs of these genes because after divergence, all the members of the gene family within zebrafish may be derived from a different original member than all of the family members in the human genome.
LG5 and LG15 have orthologs of genes on Hsa17 and duplicate copies of LHX1. Accession nos. are listed in Supplementary Data at http://www.genome.org.
Model for the Origin of Hsa17 and Its Zebrafish Orthologs
These comparative mapping data suggest one of two models for the evolution of Hsa17. The single-chromosome model suggests that the last common ancestor of tetrapods and zebrafish had a unique chromosome containing orthologs of Hsa17 genes, including multiple myhcluster genes. In the zebrafish lineage, this chromosome then broke into two unequal pieces, with some members of the myh cluster going to the smaller fragment, and others going to the larger fragment. Duplication of the larger chromosome region produced parts of LG3 and LG12, and duplication of the smaller region produced parts of LG5 and LG15. The alternative, two-chromosome model suggests that the last common ancestor of tetrapods and zebrafish had two chromosomes with orthologs on Hsa17, one with the precursors to LG3 and LG12, and the other with precursors to LG5 and LG15; these two portions would then have fused in the tetrapod lineage. This second model seems less likely because it does not easily explain the location of myh cluster genes on derivatives of both the large (LG3) and small (LG5) fragment. Mapping data from nonmammalian tetrapods should be able to distinguish these two possibilities, but the only available data are from chicken. In the chicken, the two regions orthologous to Hsa17 appear to be on different chromosomes, but the extensive fragmentation of Hsa17 orthologs into at least five chromosomes in chickens (Groenen et al. 2000) does not critically rule out either hypothesis.
Loss of Duplicated Genes
The usual fate of duplicated gene pairs is that one member is lost by mutation (Ohno 1970; Watterson 1983). What fraction of genes duplicated in the hypothesized genome-wide duplication event is present in two copies today in the zebrafish genome? Of the 74 Hsa17 loci with orthologs mapping to LG3, LG12, LG5 and LG15, at least 15 (20%) are present in duplicate. This may be an underestimate of the fraction of retained duplicates because missing duplicates may be present in the genome but not yet discovered. For loci that were duplicated in the chromosome duplication but are now known to be present as single copy on these zebrafish chromosome pairs (dlx3, hoxb2a,hoxb3a, hoxb4a, hoxb7a, hoxb10a, and eve1), the duplicate copy has either been lost from the other chromosome or has become a pseudogene by mutation (Joly et al. 1991; Ellies et al. 1997; Amores et al. 1998).
Evolution of LG3 and LG12
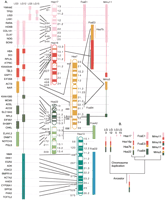
In addition to putative orthologs of Hsa17, LG3 and LG12 contain loci whose putative orthologs are from several other human chromosomes (Fig. 3). Both LGs contain several loci from Hsa16p and Hsa22. This suggests that the last preduplication ancestor of zebrafish had a single chromosome that contained orthologs of Hsa17, Hsa16p, and Hsa22. The location of apparent duplicates of the Hsa16 hemoglobin locus on LG3 and LG12 (Chan et al. 1997; Postlethwait et al. 1998) supports this conclusion.
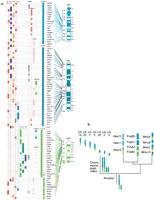
Conserved syntenies and an evolutionary model for LG3 and LG12. (A) LG3 and LG12 share conserved syntenies with Hsa16p and Hsa22 and other mammalian chromosomes in addition to Hsa17. (B) A model for the origin of Hsa17, Hsa16p, Hsa7b, and Hsa22. The overlapping conserved syntenies of orthologs of these human chromosome segments in fish, mouse, cat, and human suggest that they were originally part of a single chromosome that fragmented in the tetrapod lineage. Hsa10q and Hsa19p may have joined LG12 and LG3, respectively, after the ray-fin genome-wide duplication. Accession nos. are listed in Supplementary Data at http://www.genome.org.
Located near putative Hsa16p orthologs on LG3 are loci putatively orthologous to loci on a portion of Hsa7 called Hsa7b by Richard et al. (2000). This chromosome fragment was attached to Hsa16p until it joined the rest of Hsa7 ∼35 mya and was subsequently split in two in the human lineage by an inversion (Richard et al. 2000). Because Hsa7b orthologs are also syntenic with Hsa16p orthologs on LG3 in zebrafish, this was likely the state in the last common ancestor of zebrafish and mammals. Likewise, Mmu11 has nearly all of Hsa17, some Hsa16p, and some Hsa22 orthologs, as do LG3 and LG12, suggesting that this was the ancestral arrangement.
Taken together, these data suggest the model (Fig. 3B) that the last common ancestor of mammals and zebrafish had a single chromosome containing orthologs of loci from Hsa17, Hsa16p, Hsa7b, and Hsa22. This hypothesized large chromosome then broke apart into today's mammalian chromosomes, somewhat differently in different mammalian lineages. All or much of this large chromosome was apparently inherited intact in the zebrafish lineage. The Hsa19p and Hsa10q portions of LG3 and LG12, respectively, may have been added to, or subtracted from, ancestral chromosomes by translocation after the chromosome duplication event in the ray-fin fish lineage.
In chicken, Hsa17 orthologs are located on at least five different chromosomes, prompting the conclusion that chromosome rearrangements had joined Hsa17q25 to proximal Hsa17q in the mammalian lineage after it diverged from the avian lineage (Pitel et al. 1998). The zebrafish gene map can test that hypothesis. We have mapped the apparent orthologs of two loci from Hsa17q25; one (gdia) is on LG3 and the other (ilf1) is on LG12 (Table 1), both syntenic with many other Hsa17 orthologs. We conclude that Hsa17q25 was already syntenic with other Hsa17 loci in the last common ancestor of zebrafish and humans and separated independently in the chicken lineage after it diverged from the mammalian lineage.
The History of Hsa10q
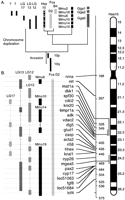
The long arm of human chromosome 10 illustrates that in some cases, the mouse genome may be more fragmented than the zebrafish genome with respect to the human genome. Numerous loci apparently orthologous to genes on Hsa10q reside on LG12 and LG13. At least three human Hsa10q genes (ADK, PAX2 and BMPR1A) are present in duplicate copies in the zebrafish genome; one copy of each is on LG12 and the other on LG13 (Pfeffer et al. 1998; Nikaido et al. 1999; Woods et al. 2000). This would be expected if these chromosomes contain duplicated segments (Fig. 4). In addition, LG17 has several putative orthologs of Hsa10q genes (Woods et al. 2000). This evidence suggests that the last common ancestor of mammals and zebrafish had a single chromosome containing Hsa10q loci. This group of genes has stayed intact in the ancestry of cats and humans but has broken up into four chromosomes in mouse.
The origin of Hsa10. (A) A model for the evolution of Hsa10. No orthologs from Hsa10p are in our data set, so the location of the zebrafish chromosome segments corresponding to Hsa10p are as yet unknown. (B) Conserved syntenies of vertebrate loci are orthologous to Hsa10q. Accession nos. are listed in Supplementary Data at http://www.genome.org.
After the duplication in the teleost lineage, fish had two orthologs of Hsa10q; one of these became part of LG12, and the other part of LG13. The Hsa10q-related portion of LG17 may have arisen by a translocation from the ancestor of either LG12 or LG13 after the chromosome duplication. Recall from Figure 1 that LG12 is largely a duplicate of LG3, but LG3 does not appear to contain a segment of Hsa10q orthologs. This suggests that the Hsa10q-related chromosome segment now on LG12 may have been either added to LG12 after the hypothesized genome duplication event or removed by translocation from LG3 after the duplication.
Hsa10p orthologs are not syntenic with Hsa10q loci in mouse or cat or zebrafish, suggesting that Hsa10q and Hsa10p joined to form today's Hsa10 after the divergence of cat and human lineages ∼90 mya (Kumar and Hedges 1998).
Hsa9
Zebrafish do not have apparent heteromorphic sex chromosomes (Endo and Ingalls 1968; Pijnacker and Ferwerda 1995; Gornung et al. 1997;Amores and Postlethwait 1999), and the genetic basis of its sex determination mechanism is as yet unknown. Hsa9 containsDMRT1, which is homologous to sex determination genes in flies and worms (Raymond et al. 1999a). DMRT1 is expressed early in the genital ridge and is in, or very near, chromosome deletions that cause sex reversal in humans (Calvari et al. 2000). Consistent with a role in sex determination, DMRT1 is on GgaZ, the sex chromosome of chickens, along with several other Hsa9 orthologs (Nanda et al. 1999). Homologues of DMRT1 are present in fish and expressed in gonads (Guan et al. 2000; Marchand et al. 2000). Thus, a zebrafish ortholog of Hsa9/GgaZ is a candidate for a cryptic sex chromosome in zebrafish.
DMRT1 and DMRT2 are closely linked in Hsa9p24.3 (Raymond et al. 1999b). We mapped terra, an apparent ortholog of DMRT2 (Meng et al. 1999) we mapped to LG5 (Kelly et al. 2000). In chicken, most Hsa9 orthologs appear on two chromosomes (Nanda et al. 1999; Groenen et al. 2000), while in mouse they reside on four chromosomes. LG5, probably the most gene-rich zebrafish chromosome, has at least 19 putative orthologs of Hsa9 genes scattered along its length, and these are distributed throughout the length of Hsa9 (Fig.5A). We conclude that the last common ancestor of birds, mammals, and ray-fin fish had a single chromosome in which all Hsa9 orthologs were syntenic. Apparently, a translocation subsequently separated orthologs of distal Hsa9q from the rest of the chromosome in the chicken lineage, and multiple chromosome fissions occurred in the mouse lineage. Thus, as with Hsa10q, more translocations appear to have disrupted the Hsa9 precursor in the mouse lineage than in the zebrafish lineage.
Antiquity of Hsa9. (A) Conserved syntenies and duplicate chromosome segments related to Hsa9. (B) A model for the origin of Hsa9. Accession nos. are listed in Supplementary Data at http://www.genome.org.
Examination of additional linkage groups suggests a history of Hsa9-related chromosome segments in zebrafish (Fig. 5B). Duplicate copies of several Hsa9 putative orthologs reside on LG5 and two other linkage groups, LG2 and LG21. These include jak2a/jak2b andnotch1a/notch1b duplicates on LG5 and LG21 andgsna/gsnb, rxrab/rxraa, and tnc/tnwduplicates on LG5 and LG2 (Woods et al. 2000). A likely model is that the duplicate of the Hsa9-related portion of LG5 broke into at least two fragments, with one portion becoming part of LG2 and the other portion becoming part of LG21. This model is testable by the mapping of more Hsa9 orthologs in zebrafish. Note that, as with Hsa17 orthologs in zebrafish, inversions have been fixed more frequently than translocations for this chromosome and that at least one inversion occurred in the human lineage since its divergence from the gorilla lineage (Hansmann 1976).
Comparative analysis of Hsa9-related chromosome segments suggests that an ancient chromosome contained orthologs of both Hsa9 and Hsa5 and that this ancient chromosome fragmented in the tetrapod lineage. The Z-chromosome of chickens consists mainly of orthologs to Hsa9 and Hsa5 (Groenen et al. 2000). Likewise, LG5 and LG21 in zebrafish have a large number of loci putatively orthologous to both Hsa9 (Fig. 5) and Hsa5 (Woods et al. 2000), and mouse chromosome Mmu13 has a contiguous block containing both Hsa5 and Hsa9 orthologs (DeBry and Seldin 1996). These data argue that this Hsa9–Hsa5 arrangement was ancestral and that in tetrapods, Hsa9 and Hsa5 separated after the divergence of the avian and mammalian lineages and in different ways in the rodent and human lineages.
Hsa11, Hsa15, and Hsa19
The orthologs of Hsa11 reside on three mouse chromosomes, Mmu2, Mmu9, and Mmu7; Hsa15 orthologs reside on the same three mouse chromosomes plus Mmu19 (Fig. 6) Mmu7 and Mmu9 also carry loci orthologous to loci on Hsa19q. In addition, chicken chromosome 5 contains a contiguous block with orthologs from Hsa11 and Hsa15 (Groenen et al. 2000). These data suggest that the last common ancestor of mouse and human had a single large chromosome that contained orthologs of Hsa11, Hsa15, and Hsa19q and that this ancestral chromosome fragmented differently in the lineages of the two mammals.
Conserved syntenies among zebrafish chromosome segments orthologous to Hsa15, Hsa11, and Hsa19. Accession nos. are listed in Supplementary Data at http://www.genome.org.
Examination of the zebrafish gene map suggests that this ancient chromosome may have existed as early as the last common ancestor of zebrafish and tetrapods. Zebrafish orthologs of Hsa11, Hsa15, and Hsa19q loci reside primarily on just a few linkage groups, and each of these has putative orthologs from all three human chromosome segments (Fig. 6A). A portion of LG25 is a duplicate of a portion of LG7 because there are at least four pairs of gene duplicates on the two chromosomes (fkd3/fkd5, isl2/isl3, pax6.1/pax6.2, andhlx1/hlx3; Woods et al. 2000). Likewise, a portion of LG18 is a duplicate of a portion of LG25 because there is at least one duplicated gene pair on these chromosomes (cyp19a/cyp19b;Chiang et al. 2000). A portion of LG5 and LG15 are duplicates because each has a copy of an LHX1 ortholog (lim1/lim6).
Because individual chromosomes in zebrafish and mouse contain putative orthologs of Hsa11, Hsa15, and Hsa19q, we conclude that these chromosomes were syntenic in the last common ancestor of zebrafish and tetrapods (Fig. 6B). After duplication in the fish lineage, losses of duplicate genes, translocations, and chromosome breaks could have produced parts of LG25, LG18, LG7, LG5, and LG15. In the tetrapod lineage, the large ancestral chromosome could have fragmented into several smaller chromosomes differently in the human and mouse lineages.
Note that, except for EFNA2, this set of zebrafish linkage groups does not contain putative orthologs of loci on the short arm of Hsa19. LG22, LG3, and LG2, however, each has several Hsa19p loci (Fig.6A, Rubenstein et al. 2000). This suggests that Hsa19p and Hsa19q were not syntenic in the last common ancestor of zebrafish and human. This conclusion is supported by the genetic map of the cat, in which FcaA2 is orthologous to Hsa19p and FcaE2 is orthologous to Hsa19q (Murphy et al. 2000). Hsa19p and Hsa19q apparently joined after the divergence of cat and human lineages. Three other linkages that are ancestral in mammals, Hsa19q and Hsa16p, Hsa14 and Hsa15, and Hsa12 and Hsa22 (Chowdhary et al. 1998; O'Brien and Stanyon 1999; Murphy et al. 1999;Richard et al. 2000), are not apparent in the current zebrafish data set, suggesting that these chromosome segments may have joined in the tetrapod lineage before the mammalian radiation.
DISCUSSION
The results presented here address some of the mechanisms by which humans and other tetrapods arrived at their chromosome numbers and configurations. First, the results test whether a mechanism of karyotypic evolution proposed for mammals, the prevalence of translocations over inversions (Ehrlich et al. 1997), holds for vertebrates in general. If this hypothesis reflects a general mechanism, then blocks of conserved synteny should be small, but gene order should be retained within blocks. Alternatively, if inversions are more prevalent than translocations, blocks of conserved synteny should be large with gene orders more scrambled.
Comparative analysis shows first that blocks of conserved synteny are large between zebrafish and humans. Syntenic groups the size of some of today's human chromosomes and long chromosome arms existed >450 mya in the last common ancestor of zebrafish and humans. Examples include LG3 and most of Hsa17 (Fig. 1); LG12 and Hsa10q (Fig. 4); LG5 and all of Hsa9 (Fig. 5), which is broken into at least five chromosomes in chicken (Groenen et al. 2000) and four in mouse; LG22 and Hsa19p (Fig.6), which joined Hsa19q after the divergence of carnivore and primate lineages; LG17 and Hsa14 (Woods et al. 2000); LG9 and Hsa2q (Postlethwait et al. 1998; Amores et al. 1998; Hsa2q joined Hsa2p after the divergence of human and chimpanzees lineages [Weinberg et al. 1994]); LG8 and the short arm of Hsa1 (Woods et al. 2000; Hsa1p joined Hsa1q within the primate lineage [Richard et al. 1996]); and LG19 and Hsa6p, the environs of the major histocompatibility locus (Bingulac-Popovic et al. 1997; Michalova et al. 2000). This evidence shows that translocations have not disrupted many chromosome segments that were present in the last common ancestor of zebrafish and humans. Zebrafish, however, also has many chromosome segments orthologous to smaller portions of human chromosomes, showing that despite some large conserved regions, translocations have also disrupted syntenies in the two lineages.
Although some lengthy chromosome segments are conserved between zebrafish and human, the orders of loci within chromosome segments are substantially rearranged. This is demonstrated here for LG3/Hsa17 and LG5/Hsa9. This suggests that chromosome inversions have frequently been fixed in diverging populations in the lineage leading to zebrafish. Genomic DNA sequencing in the pufferfish Fugu rubripes has shown that even over a stretch of a dozen or so genes, transpositions and inversions have sometimes rearranged fish chromosomes with respect to humans by a breakpoint about every megabase or so in human (Brunner et al. 1999; Elgar et al. 1999). Thus, transpositions of locus order within large conserved blocks of synteny appear to support the hypothesis that inversions have been a more frequent force in the shaping of vertebrate karyotypes than translocations, although both processes have clearly played a role.
A comparison of duplicated chromosome segments can reveal the dynamics of chromosome rearrangements since the duplication event. Because the orders of duplicated genes on LG3 and LG12 are not colinear, inversions have occurred in the fish lineage after the genome duplication event. Likewise, inversions have occurred in the mouse and human lineages altering gene orders in the orthologous portions of Hsa17 and Mmu11 (see http://www.ncbi.nlm.nih.gov/Homology/Davis/). Translocations have also occurred in the fish lineage after the genome duplication. Evidence for this conclusion comes from LG3, LG12, and LG13: A portion of LG12 is orthologous to Hsa10q, but the duplicate copy of Hsa10q is on LG13, not LG3, which otherwise is the duplicate of LG12. Thus, Hsa10q-related loci were either translocated to LG12, or away from LG3, after the chromosome duplication.
The analysis presented here allows a preliminary estimate of the frequency of gene duplicates that have been retained since the genome duplication in the fish lineage. Of the 74 Hsa17 loci with putative orthologs on LG3, LG5, LG12, and LG15, at least 15 (20%) are present in duplicate copies. Likewise, LG2, LG5, and LG21 have putative orthologs of 22 Hsa9 loci, with five (23%) present in duplicate. Because some loci mapped to these chromosomes may have duplicates that remain to be discovered, the observed fraction of preserved duplicates may be an underestimate of the true value. Thus, 20% or more of the genes duplicated in the presumed genome duplication event may still survive. Experiments suggest that the preserved zebrafish duplicates are no longer totally redundant in function but may have partitioned ancestral functions between the two surviving gene copies or evolved novel functions (for discussion, see Force et al. 1999).
In addition to instances for which zebrafish has two orthologs of mammalian genes, zebrafish has, in other cases, additional genes for which the mammalian copy has been lost. For example, members of theEVX gene family lie adjacent to the 5′ ends of theHOXA and HOXD clusters in zebrafish and humans, but zebrafish has, in addition, eve1 at the homologous location adjacent to a zebrafish ortholog of the HOXB cluster (Joly et al. 1991; Amores et al. 1998). The last common ancestor of zebrafish and human must have had an evx gene adjacent to itshoxb cluster, but that gene apparently became a pseudogene in the mammalian, but not the zebrafish, lineage. Similarly,hoxc1 and hoxc3 are present in fish (Aparicio et al. 1997; Amores et al. 1998; Naruse et al. 2000), but the orthologs are missing from mammals, showing that ancestral gnathostomes had a greaterhox repertoire than extant tetrapods. The ependymingene (Sterrer et al. 1990) on LG5 may represent a similar case.
The complementary loss of closely linked gene duplicates can explain genome arrangements that are otherwise difficult to understand. For example, class I and class II loci of the major histocompatibility complex are unlinked in zebrafish (Postlethwait et al. 1994;Bingulac-Popovic et al. 1997; Michalova et al. 2000), but they are tightly linked in mammals. A hypothesis to explain this situation assumes that class I and class II genes initially arose by tandem duplication and that this was the condition in the last common ancestor of zebrafish and humans. Immediately after the ray-fin fish genome duplication, there were two chromosomes, each with redundant, closely linked class I and class II loci and surrounding genes. If the class I loci were lost on one of the duplicates, the class II loci were lost on the other duplicate, and other loci similarly shared complementarily between the two regions, then the present day situation in teleosts and mammals could be explained by purely degenerative processes without having to resort to postulating precise translocations to construct today's MHC arrangements.
If the multiple copies of zebrafish chromosome segments arose by whole genome duplication (Amores et al. 1998; Postlethwait et al. 1998; Gates et al. 1999; Woods et al. 2000), zebrafish should have twice as many chromosomes as humans in the absence of chromosome rearrangements. But zebrafish, with 25 chromosomes, has just two more chromosomes in the haploid set than humans. Two general hypotheses might account for this situation. In the fish fusion model, the last common ancestor of zebrafish and humans had ∼24 chromosomes, and an excess of chromosome fusions in the fish lineage before or after the genome duplication reestablished the original number. Alternatively, in the tetrapod fission model, the ancestral state was ∼12 chromosomes, which doubled in the fish lineage but fragmented in different places in different tetrapod lineages to establish current karyotypes.
Both models predict that zebrafish chromosomes will be mosaics of tetrapod chromosomes, but they make different predictions regarding the relationships of those chromosome fragments. In the fish fusion model, zebrafish chromosomes should have no regular relationship to arrangements of various tetrapod chromosomes. Alternatively, the tetrapod fission model predicts that zebrafish should have syntenic sets of genes that are found to be broken up differently in chromosomes of different tetrapod lineages, so that tetrapods would have overlapping subsets of orthologs that are syntenic in zebrafish.
The comparative analysis in Figures 3 and 6 suggest the following conclusions: Among the chromosomes of the last common ancestor of teleosts and mammals, one chromosome contained the precursor of all or major parts of Hsa17, a small portion of Hsa7 (Hsa7b of Richard et al. [2000]), Hsa16p, and Hsa22, and another chromosome contained the ancestors of major portions of Hsa11, Hsa15, and Hsa19q. The model assumes that these chromosome segments remained syntenic in the lineage leading to zebrafish and persisted after the genome-wide duplication, with pruning of most duplicate genes, but also assumes that these chromosomes were broken up differentially in different tetrapod lineages. The overlapping conserved syntenies found in rodents and human, which may have diverged rather early in the radiation of eutherian mammals (Kumar and Hedges 1998; Mindell et al. 1999;Penny et al. 1999; but see Gatesy et al. 1999; Waddell et al. 1999; Liu and Miyamoto 1999), provide evidence for this model. For example, Mmu7, rat chromosome Rno1 (Watanabe et al. 1999), and zebrafish LG18, LG25, LG7, and LG5 are all orthologous to portions of Hsa11, Hsa15, and Hsa19q, and chicken chromosome Gga5 contains portions of Hsa11 and Hsa15 orthologs (Groenen et al. 2000), suggesting that the ancestral chromosome contained this entire set of genes and that after intrachromosomal rearrangements, it broke in different ways in the different tetrapod lineages.
Loci that are closely linked in zebrafish and in one tetrapod, suggesting that they were closely linked in the last common ancestor, can be on two or three chromosomes in a different tetrapod. For example, orthologs of APOA (apoa, Y13653),HEXA (hexa, AI629215), and ATM(atm, AI721600) span a 3-CM interval on Mmu9 and 15 cM on zebrafish LG5 but are on two chromosomes (Hsa11 and Hsa15) in human; similarly, the orthologs of CCNE1 (ccne, X83594),IGF1R (igf1r, AI330927), and EIF4G2(eif4g2, AI584683) are in a 35-CM stretch of Mmu7 and are syntenic on LG7 but are on three chromosomes in human (Woods et al. 2000). In other cases, for example, on Hsa9/LG5 and Hsa10/LG12, zebrafish and human share syntenies, and the orthologs are on multiple chromosomes in mouse. In these instances, the zebrafish genome acts as an outgroup, and thus suggests the primitive chromosomal condition, while one or the other of the mammalian genomes represents a more recently derived state, the differential fission of the ancestral chromosome.
Although the mapping data suggest that, on average, mammalian genomes derive from the fission of large ancestral chromosomes, clearly chromosome fusions have also occurred in the human lineage. For example, Hsa1p and Hsa1q fused within the primate lineages (Dutrillaux et al. 1978), and Hsa2p and 2q fused after the divergence of chimpanzee and human lineages (Weinberg et al. 1994).
If the tetrapod fission model is correct, then the last common ancestor of zebrafish and humans may have had something in the range of 12 or 13 chromosomes in the haploid set. These would then have doubled to the 24 or so present in most teleosts and would have broken apart into the 20–30 present in many Eutherian mammals. Independent support for this model comes from the ancestral Metatherian (marsupial) karyotype, which consisted of ∼7 chromosomes (DeLeo et al. 1999), consistent with the suggestion that the last common ancestor of marsupial and placental mammals may have had fewer chromosomes than most of today's Eutherian mammals.
The comparative mapping reported here has analyzed in detail the histories of some duplicated chromosome segments in zebrafish and investigated the evolution of mammalian chromosomes using zebrafish as an outgroup to help infer the ancestral state of the karyotype. Further work needs to focus on the consequences of the ray-fin fish genome-wide duplication event. When did the event documented in zebrafish occur in fish phylogeny? Did it occur at the base of teleost radiation, as the current scanty evidence suggests, or even earlier in the ray-fin lineage, or later in several teleost lineages independently? (Genome duplications in salmonids and carp have almost surely occurred after, and on top of, the genome-wide duplication detected in zebrafish [Allendorf and Thorgaard 1984; Larhammar and Risinger 1994].) What fraction of duplicate genes have been retained in zebrafish, and is this reflective of ray-fin fish genomes as a whole? Are different sets of duplicate genes retained in different lineages of ray-fin fish? If different genes remain duplicated in different teleosts, then how do the functions of duplicates in one species compare to those of singletons in a related species? What do these between-teleost comparisons tell us about the role of duplicated genes in speciation and the evolution of novel features? The model of chromosome fission proposed here for the ancestry of Eutherian chromosomes must be further tested by detailed gene mapping in species providing information on important phylogenetic nodes; especially valuable species would include a Metatherian such as the wallaby (Wilcox et al. 1996), the chicken (Groenen et al. 2000), and a frog (Amaya et al. 1998). It will be revealing to examine these gene maps in light of a fully sequenced human genome.
METHODS
Sequences of Danio rerio genes and ESTs (M. Clark and S. Johnson, Washington University Zebrafish Genome Resources Project;http://zfish.wustl.edu) were obtained from NCBI, and primers were designed as described by Kelly et al. (2000). SSLP (simplesequence length polymorphism) primer sequences were obtained from Shimoda et al. (1999) andhttp://zebrafish.mgh.harvard.edu/. Primers were synthesized at the Stanford Genome Technology Center or obtained from Research Genetics. Polymorphisms were detected and mapped in the heat shock mapping panel as described (Kelly et al. 2000). The genotype data set for the markers mapped in this article are available athttp://www.neuro.uoregon.edu/postle/mydoc.html, and a complete data set of all markers on the HS panel is found athttp://zebrafish.stanford.edu. Maps were constructed with MapManager (Manly 1993).
Zebrafish genes and ESTs were assigned putative human orthologs by BLASTX searches (Altschul et al. 1997) of zebrafish nucleotide sequences against the NCBI human nonredundant protein sequence database (http://www.ncbi.nlm.nih.gov/blast/blast.cgi). Where possible, we concatenated 5′ and 3′ sequences of ESTs for BLASTX searches. When the results of these searches had expect scores (Evalues) of 1e−5 or less, the putative orthologs were further tested with reciprocal searches against the zebrafish subset of nonredundant sequences (NR) and dbEST databases. A human ortholog was confirmed if the original zebrafish gene or EST (or a gene or EST in the same UniGene cluster) was in the top five matches of the reciprocal search by TBLASTN. Woods et al. (2000) tested the validity of this procedure by comparing putative orthologs assigned from 43 different cloned genes and from 95 ESTs derived from those genes. They found no cases of different ortholog assignments, showing that this method works well, although we realize that for some gene families, mistakes will sometimes happen. Map positions for mammalian orthologs were found using the OMIM (http://www.ncbi.nlm.nih.gov/Omim), LocusLink (http://www.ncbi.nlm.nih.gov/LocusLink), and GeneMap'99 (http://www.ncbi.nlm.nih.gov/genemap99) databases and occasionally by a BLASTN search of the human nucleotide sequence against the htgs (high throughput genomesequencing) database at NCBI. Mouse orthologs were identified using the HomoloGene database (http://www.ncbi.nlm.nih.gov/HomoloGene), and their map locations were found using Locuslink and the Mouse Genome Database (http://www.informatics.jax.org). Orthologs and their map positions are listed in Table 1.
Either because the putative human orthologs of some of the zebrafish genes we mapped had not been mapped or because their positions were not known precisely enough to infer relative orders, we placed 17 human genes on the G3 radiation hybrid mapping panel following standard protocols (http://www-shgc.stanford.edu/Mapping/index.html). The vectors and full results are available athttp://www.neuro.uoregon.edu/postle/mydoc.html.
Acknowledgments
We thank Tim Cardozo, Tom Conlin, and Allen Day for expert help in bioinformatics; discussions with Allan Force, Angel Amores, and Tom Titus, and the Stanford Genome Technology Center for oligonucleotide synthesis. This work was supported by NIH grants R01DK55378 (W.S.T. and J.H.P.), R01RR12349 (W.S.T.), and R01RR10715 and NSF grant IBN-9728587 (J.H.P.). We thank the National Institutes of Health (1-G20-RR11724), National Science Foundation (STI-9602828), M.J. Murdock Charitable Trust (96127:JVZ:02/27/97), and W.M. Keck Foundation (961582) for supporting renovation of the University of Oregon Zebrafish Facility. W.S.T. is a Pew Scholar in the Biomedical Sciences.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
-
↵3 Corresponding author.
-
E-MAIL jpostle{at}oregon.uoregon.edu; FAX (541) 346-4538.
-
Article and publication are at www.genome.org/cgi/doi/10.1101/gr.164800.
-
- Received September 13, 2000.
- Accepted October 24, 2000.
- Cold Spring Harbor Laboratory Press