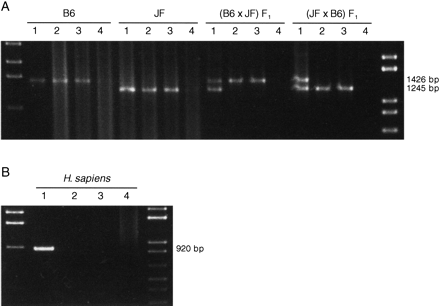
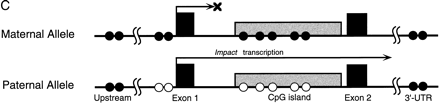
Parent-of-origin-specific methylation of Impact. (A) Methylation-specific PCR assays for the CpG island of mouseImpact. PCR products from native genomic DNA and those digested with HhaI, HpaII, or MspI (lanes1–4, respectively) were subjected to 1.5% agarose gel electrophoresis and stained with ethidium bromide. StyI-digest λ DNA and 1 kb PLUS DNA LADDER (GIBCO BRL) were used as size standards in the left- and rightmost lanes, respectively. The mouse intronic CpG island was analyzed in JF, B6 (B6 × JF) F1, and (JF × B6) F1. (B) Methylation-specific PCR assays for the CpG island of human IMPACT. The human CpG island, which overlaps the promoter region and lacks length polymorphisms, was also analyzed as described in A. (C) Model for the imprinted expression of mouseImpact. Exons are depicted as solid boxes, and CpG islands are shaded. The island is a region of differential methylation, where only the maternal allele is hypermethylated. Closed and open circles stand for hypermethylation and undermethylation, respectively.











