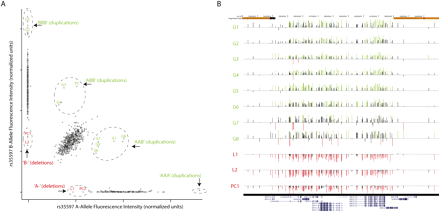
CNVs at 16p13.11 in individuals with ID. (A) SCOUT uses fluorescence intensity data generated for SNP genotypes, in this case at rs35597, to identify samples that harbor deletions or duplications. Each sample is assigned a score based on its intensity relative to the center of the cluster of all samples with the same genotype. Samples that harbor duplications exhibit greater total intensity and potentially aberrant allelic ratio (inferred genotypes marked in green), while deletions exhibit lesser intensity and lack SNP heterozygosity (inferred genotypes labeled in red). Samples analyzed as positive controls are labeled PC, while validated predictions are labeled as L1–L2 (“loss”) or G1–G9 (“gain”). Dashed circles are drawn manually for visual aid. Unlabeled samples that appear to be outliers at this probe are either outliers only at this probe or samples with insufficient DNA for validation. (B) SCOUT predictions were validated with targeted array-CGH experiments. Shown are the resulting data for the labeled samples in panel A, with normalized log-transformed test/reference ratios at each tested probe presented as vertical bars. Those probes that are more than 1.5 standard deviations above (below) the average for the entire array are colored green (red). The samples shown include eight validated duplications, two validated deletions, and one positive control deletion. Note that G9 was analyzed with a different array-CGH platform and is not shown here. Genome coordinates and segmental duplications are annotated at the top. The bottom shows RefSeq genes/isoforms annotated in this region.











