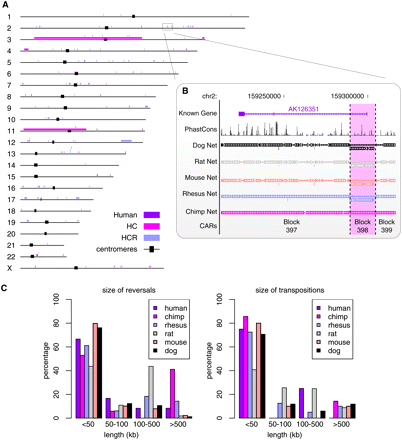
Localization on the human genome of the primate reversals and length distribution of the predicted rearrangement events. (A) Reversals recovered on the path from the primate-rodent ancestor to the human. These reversals are human–chimp–rhesus (HCR) specific, human–chimp (HC) specific, or human specific. Some reversals are displayed at different heights if they are too close (e.g., the three small regions in Chr13). (B, highlighted in pink) Example of an HC-specific reversal recovered by EMRAE. The reversal is shown on human chromosome 2 along with the UCSC net tracks for the other genomes. Synteny block (or CARs) 398 has opposite orientation in rhesus, mouse, rat, and dog as compared to human and chimp. (C) Length distribution of the reversed and of the transposed regions for the events predicted for the extant genomes.











