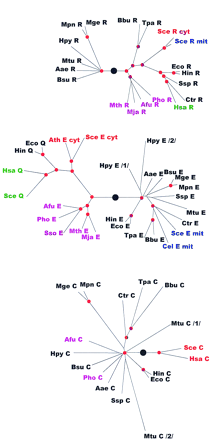
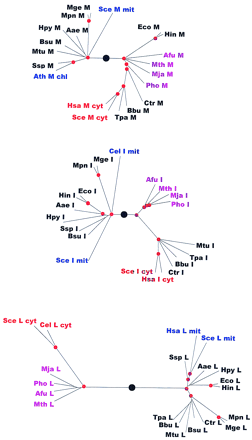
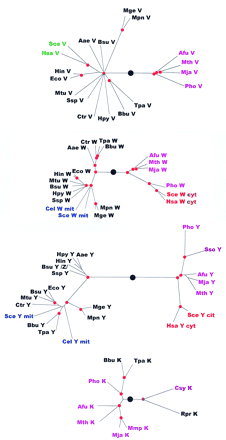
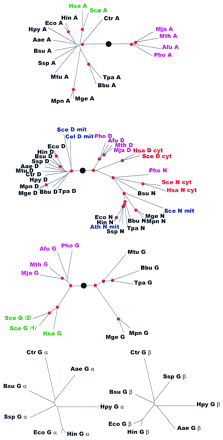
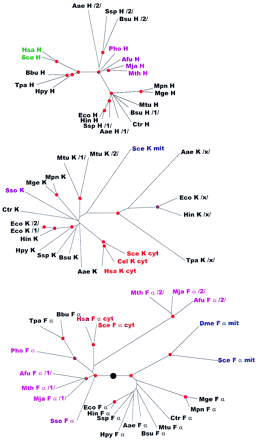
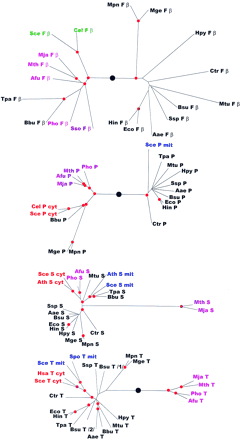
Phylogenetic trees for aaRSs. The trees are shown as unrooted, but a large black circle denotes the likely root position inferred by a combination of synapomorphy analysis, clustering by sequence similarity and midpoint rooting (see Table 2). Red circles mark statistically supported nodes (> 70% bootstrap support for both the neighbor-joining and the Fitch–Margoliash methods). Three-letter species labels are as in Fig. 1. Labels for archaeal proteins are shown in magenta; eukaryotic cytoplasmic in red; eukaryotic organellar in blue; eukaryotic without indication of origin in green; bacterial in black. Additional abbreviation: (Sso) S. solfataricus.











