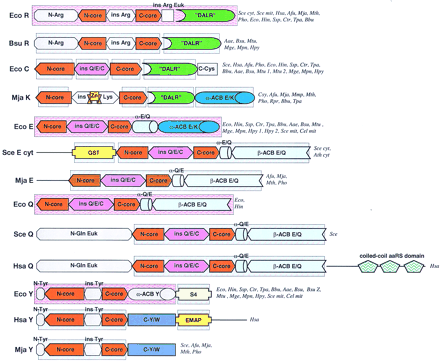
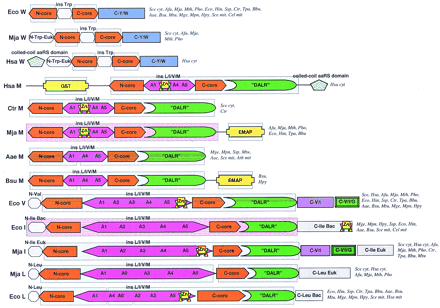
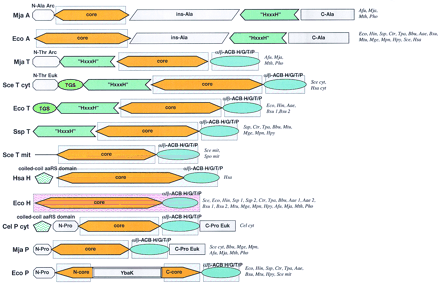
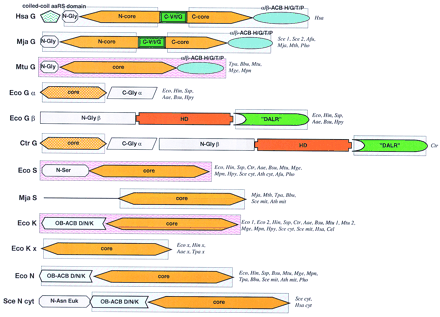
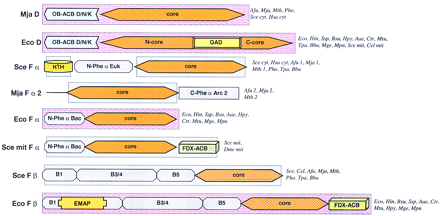
Domain architectures of aaRSs. (A) Class I aaRSs; (B) class II aaRSs. For each specificity, all detected distinct domain architectures are shown. The abbreviated names of the species names in which the given domain arrangement was observed are given to the right of each scheme. Domain name abbreviations: (A1–5) Distinct modules detectable in the large insert typical of the aaRS for aliphatic amino acids; (ACB) anticodon-binding domain; (FDX–ACB) ferredoxin-fold ACB; (GST) glutathione S-transferase; (ins) insert; (OB–ACB) OB-fold ACB (Zn) Zn ribbon motif; other domains are designated either after partially conserved sequence signatures (DALR and HxxxH) or after the aaRS specificities in which they are found (C-V/I, C-V/I/G). Pink background indicates domains for which three-dimensional structure is available; blue background shows domains for which the structure can be predicted on the basis of sequence similarity to structurally characterized domains. (Aae) A. aeolicus; (Afu) A. fulgidus; (Ath) A. thaliana; (Bbu) B. burgdorferi; (Bsu) B. subtilis; (Cel) C. elegans; (Csy) C. symbiosum; (Dme) D. melanogaster; (Hsa)H. sapiens; (Ctr) C. trachomatis; (Eco) E. coli; (Hin) H. influenzae; (Hpy) H. pylori; (Mja)M. jannaschii; (Mge) M. genitalium; (Mpn) M. pneumoniae; (Mth) M. thermoautotrophicum; (Mtu) M. tuberculosis; (Mpn) M. pneumoniae; (Pho) P. horikoshii; (Rpr) R. prowazekii; (Sce) S. cerevisiae; (Spo) S. pombe; (Ssp) Synechocystissp. (Tpa) T. pallidum.











