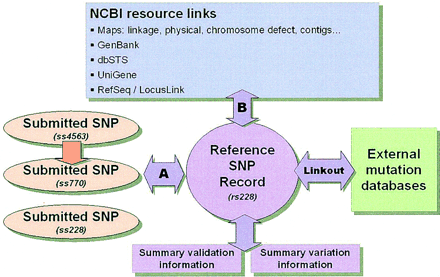
The NCBI dbSNP data model. Individual submissions to dbSNP are represented by the three ovals on the left. Postsubmission computations identify sets of identical submissions and cluster them into single reference SNP records (step A). Submissions that are determined to be unique at the time of analysis also have reference SNP records constructed at this time as well. All reference SNP records are analyzed by a second round of BLAST analysis in an attempt to annotate them onto other NCBI resources (step B) as discussed in the text. Illustrated here is a real case in the database in which two groups independently reported a variation (ss228, ss770), and a third group (ss4563) submitted additional frequency data on the published variation ss770. All three records were grouped into the single reference record, rs228. Links are established between individual submissions and external databases or servers, such as locus-specific mutation databases or submitter web sites by using the LINKOUT line type. These links will direct users to additional information on a particular variation.











