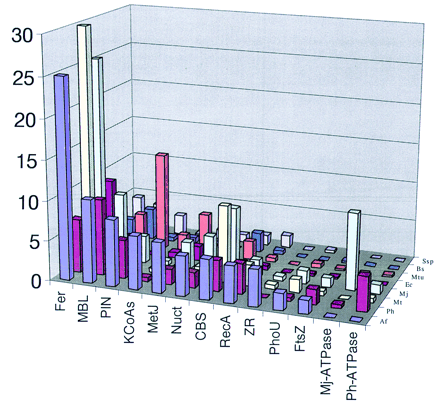
Specidic expansion of protein families in Euryarchaeota. The members of the families were identified using family-specific PSSMs are described in Methods. (Vertical axis) Number of proteins (domains) per 1000 genes. (Fer) Ferredoxins; (MBL) metallo-β-lactamase; (Nuct) “minimal” nucleotidyltransferase (Koonin et al. 1997; Aravind and Koonin 1999b); (PIN) PilT-amino-terminal domain (see text); (CBS) cystathionine-β-synthase domain; (FtsZ) GTPases involved in cell division, orthologs of the bacterial FtsZ protein; (RecA) superfamily ATPases; (MetJ) Arc/Met-repressor class of transcription regulators; (PhoU) regulators of phosphate uptake, orthologs of the bacterial PhoU protein; (KCoAS) ketoacyl-coenzyme A synthetases; (ZR) a distinct, archaea-specific family of predicted nucleic acid-binding protein containing the zinc ribbon domain (L. Aravind, unpubl.). Archaea; (Af)A. fulgidus; (Mj) M. jannaschii; (Mta) M. thermoautotrophicum; (Ph) P. horikoshii. Bacteria: (Bs)Bacillus subtilis; (Ec) Escherichia coli; (Mtu)Mycobacterium tuberculosis; (Ssp) Synechocystis sp.











