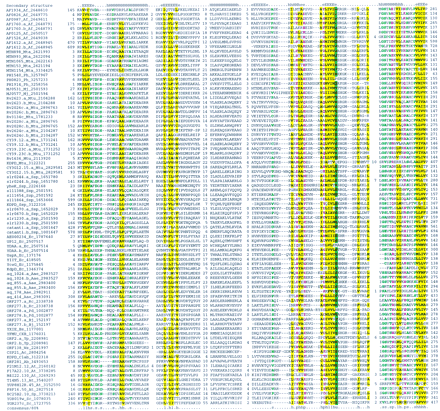
Previously undetected protein family conservation in archaea, bacteria, and eukaryotes—the UspA superfamily of predicted nucleotide-binding, regulatory proteins. The alignment was constructed on the basis of the PSI-BLAST results using the Clustal W program. The inclusion of each sequence in the superfamily was statistically supported by the PSI-BLAST analysis with an e-value of at least 0.01. The left column includes the protein (gene) names, and the gene identification (GI) numbers (after the underscore). A consensus derived using the 80% cutoff is shown underneath the alignment and the respective alignment columns are highlighted; (b) a “big” residue (E,K,R,I,L,M,F,Y,W); (h) hydrophobic residues (A,C,F,I,L,M,V,W,Y); (s) small residues (A,C,S,T,D,N,V,G,P); (u) “tiny” residues (G,A,S); (p) polar residues (D,E,H,K,N,Q,R,S,T); (c) charged residues (K,R,D,E,H); (_) negatively charged residues (D,E). The distances from the aligned regions to the protein termini and the distances between the conserved blocks, where more variable regions were omitted, are indicated by numbers. The secondary structure elements predicted using the PHD program, with the multiple alignment as the input (Rost and Sander 1994), is shown above the alignment; (E) extended conformation (β-strand); (H) α-helix. Species name abbreviations: (AaeAquifex aeolicus; (Ab) Azospirillum brasilense; (Ac)Acanthamoeba castellanii; (Af) Archaeoglobus fulgidus; (At) Arabidopsis thaliana; (Bj)Bradyrhizobium japonicum; (Bs) Bacillus subtilis; (Clab)Clostridium acetobutylicum; (Cxb) Coxiella burnetti; (Ec) Escherichia coli: (Hi) Haemophilus influenzae; (Mj) Methanococcus jannaschii; (Mta) Methanobacterium thermoautotrophicum; (Mtu) Mycobacterium tuberculosis; (Ph) Pyrococcus horikoshii; (Pd) Paracoccus denitrificans; (Rc) Rhodobacter capsulatus; (Sp) Schizosaccharomyces pombe; (Ssp) Synechocystis sp.











