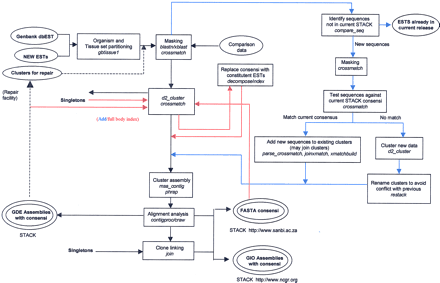
STACK processing overview. Inputs are shown in single-line ellipses, outputs in double-line ellipses. STACK first iteration, ADD, INDEX phases, and the repair facility are indicated by black, blue, red, and black-dotted arrows respectively. In the first iteration (black arrows), human sequences from GenBank dbEST are partitioned into manageable, tissue-related sets. Common vector and repeat sequences are masked, and the resulting entries are subjected to loose clustering by d2_cluster. Clusters of related sequences are assembled by PHRAP, and their alignments are analyzed by CRAW. GDE format assembly data are output, and CONTIGPROC selects appropriate consensi and subconsensi. Available clone-ID information is used to identify clone-linked clusters, after which full-length, joined consensus sequences are output in FASTA (Pearson) format. Complete assembly and linkage information is saved for each index class in GIO format (NCGR). ADD (blue arrows) incorporates new sequence data by comparison to existing STACK consensi. Existing clusters that are identified as members of the same group are reassembled and submitted to the STACK_PACK processing pipeline in combination with and newly generated D2 clusters. During the whole body index (red arrows), all cluster consensi and singletons are submitted as a single set to D2_ cluster during the whole-body index phase (red arrows). The resulting index clusters are then expanded prior to assembly by replacing each consensus with the sequences that contribute to it.











