Functional Genomics and Rat Models
The 20th Century has seen a remarkable number of inventions and technological advances in virtually all aspects of human life and health care. Many areas of biomedical research have made great strides in unraveling the cause of human disease and in developing new therapies to counter, or at least improve, outcome from disease. However, the cause of the vast majority of common disease remains poorly defined. In the final year of the millennium, the release of the draft sequence of the human genome promises to bring in a new era for basic science research and, hopefully, unprecedented growth in our understanding of human disease. For this to occur there is a critical need to annotate the genomic sequence with gene function and basic biology. Typically, the view from the geneticist immediately turns to mouse, as the mammalian contributor. Yet, not all biologists are willing to convert to the mouse as their system of choice, in many cases because of the existence of better models. Although the mouse is undoubtedly going to play a major role in contributing to the annotation of gene function, other mammalian species will also make significant contributions. This Insight/Outlook piece focuses on the role the rat will play in annotating the genome in the functional genomics era.
The laboratory rat, Rattus norvegicus, was the first mammalian species domesticated for scientific research, with work dating back to before 1850 (Lindsey 1979). From this auspicious beginning, the rat has become the most widely studied experimental animal model for biomedical research. Since 1966 (the earliest year covered by the Medline database), nearly 500,000 research articles reporting the use of rats have been published, most focused on evaluating the biology and/or the pathobiology of the rat. In contrast to its central role in the study of behavior, biochemistry, neurobiology, physiology, and pharmacology, the rat has lagged far behind the mouse as a genetic “model” organism, until recently.
Historically, rat genetics had a surprisingly early start. The first genetic studies were carried out by Crampe from 1877 to 1885 and focused on the inheritance of coat color (Lindsey 1979). Hugo De Vries, Karl Correns, and Erich Tschermak rediscovered Mendel's laws at the turn of the century, and Bateson used these concepts in 1903 to demonstrate that rat coat color is a Mendelian trait (Lindsey 1979). The first rat inbred strain, PA, was established by King in 1909—the same year that inbreeding began for the first inbred strain of mouse, DBA (Lindsey 1979). Despite this parallel start, the mouse soon became the model of choice for mammalian geneticists, whereas the rat became the model of choice for physiologists, nutritionists, and other biomedical researchers. Geneticists preferred the mouse because of its smaller size, which simplified housing requirements, and the availability of many coat color and other mutants exhibiting Mendelian patterns of inheritance, which had been collected by mouse fanciers (Nishioka 1995). Physiologists and other biomedical researchers favored the rat because its larger size facilitated experimental interventions. Over time a large number of rat strains were used to develop disease models by selective breeding, which “fixes” natural disease alleles in particular strains or colonies (Greenhouse et al. 1990). For example, there are inbred strains of rats used for research in the following areas: addiction, aging, anatomy, autoimmune diseases, behavior, blood diseases, breast cancer, cardiovascular diseases, cancer, comparative genomics, dental diseases, diseases of the skin and hair, endocrinology, eye disorders, growth and reproduction, hematologic disorders, histology, kidney diseases, metabolic disorders, neurological and neuromuscular diseases, nutrition, pathophysiology, pharmacology, pulmonary diseases, physiology, reproductive disorders, skeletal disorders, sleep apnea, transplantation and immunogenetics, toxicology, and urological disorders (Gill et al. 1989; Greenhouse et al. 1990; James and Lindpaintner 1997).
Why Rats?
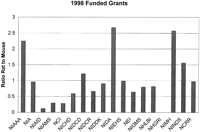
With all the success in the mouse genome project, including the initiation of the mouse genomic sequencing project and the prospect of having every mouse gene knocked out, why should there be interest in rat? Although there are many ways to answer this question, the fundamental reason is the biology. However, I think it is important to illustrate how much research is conducted in rats. Most investigators are aware that the rat remains a major model system inside the pharmaceutical industry; yet most are surprised as to how much research is conducted in rats inside academia. One of the major reasons for this surprise is the lack of a vocal rat community. Investigators that use rats tend to be disease based and not model based. For example, in my own research, I have worked in rat, mouse, zebrafish, and human. Inside the National Institutes of Health (NIH) extramural program, the number of research grants funded using rats on average nears parity for most institutes and centers (Fig. 1). Finally, comparing a Medline search on rat versus mouse reveals that rat research is much more mechanistic in content (data not shown). Therefore, it seems logical to position the rat field so the mechanistic, disease-based research can be integrated into the awesome power of the human and mouse genome projects.
The relative ratio of grants funded for research using rat and mouse. The results come from searching CRISP (ComputerRetrieval of Information onScientific Projects) reports from the NIH for fiscal year 1998. For a variety of reasons CRISP reports are not very accurate; so a ratio between mouse grants and rat grants was used. In this way, the inaccuracies of the CRISP reports are hopefully at least consistent. A ratio of 1 means the same number of grants is funded within the institute or center. A ratio >1 indicates more rat grants are funded, and a ratio <1 indicates more mouse grants are funded. NHGRI does not show as funding rat. This is an example of the inaccuracies of CRISP, as NHGRI has contributed resources to the rat genome project. Abbreviations: National Institute on Alcohol Abuse and Alcoholism (NIAAA), National Institute on Aging (NIA), National Institute of Allergy and Infectious Diseases (NIAID), National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), National Cancer Institute (NCI), National Institute of Child Health and Human Development (NICHD), National Institute on Deafness and Other Communication Disorders (NIDCD), National Institute of Dental and Craniofacial Research (NIDCR), National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK), National Institute on Drug Abuse (NIDA), National Institute of Environmental Health Sciences (NIEHS), National Eye Institute (NEI), National Institute of General Medical Sciences (NIGMS), National Heart, Lung, and Blood Institute (NHLBI), National Human Genome Research Institute (NHGRI), National Institute of Mental Health (NIMH), and National Institute of Neurological Disorders and Stroke (NINDS), National Center for Research Resources (NCRR).
Progress of the Rat Genome Project
Recognizing the usefulness of the rat as a model system, NIH, led by the National Heart, Lung, and Blood Institute (NHLBI), has funded the Rat Genome Project (RGP), the Rat Expressed Sequence Tag (RGPEST) Project, and the Rat Genome Database (RGD) to develop important genomic tools and resources that will further enhance the power of rat model systems (Table 1). However, it is important to point out that this effort has been an international effort, albeit a relatively small group of 13 labs that I know about (Rat Genome Groups). This international team has now built the beginnings of a rat “backbone map” using the rat radiation hybrid (RH) panel. This backbone map contains the vast majority of genetic markers from the major laboratories that developed the markers on a single RH map. Rat is the first organism with a dense, single integrated map, yielding a means to integrate all previously mapped genes and quantitative trait loci onto a single backbone. This backbone map forms the foundation of the recently U.S. sponsored RGD, which is coordinated with RATMAP, the Mouse Genome Database (MGD), and the National Center for Biotechnology Information (NCBI).
Rat Genome Resources
The development of genomic resources for the rat has been enhanced by the U.S. RGPEST project that has just been renewed and whose 3-year goals are to identify 60,000+ unique rat genes (UniGenes) and to map over half of them using the rat RH panel, as well as the funding of the RGD. In addition the German Genome Project has just been funded for the construction of a physical map for the rat. Notably, the U.S. and German projects continue to coordinate their activities. Consequently, the infrastructure for the rat is rapidly reaching a level whereby sequencing the genome is the next logical choice.
To Sequence or Not To Sequence
Given that the mouse genomic sequencing project has started with the primary purpose to provide a means to annotate the human genome with function. What is the rationale for sequencing the rat genome project anytime soon? The answer is in the comparative genomics that can be conducted with three mammalian species versus two. First, the rat and the mouse have been evolutionary separated for ∼16 million years, whereas human and a common rat–mouse ancestor have been split for ∼80 million years. These estimates are derived using the equation r = K /(2T), in which ris the rate of mutation, K is the distance, and Tis the divergence time between the two species (Li 1997) and the estimates of K and r from Makalowski and Boguski (1998). They also conclude that human and rodent may be too distant to detect subtle differences in substitution distances within murine (Makalowski and Boguski 1998), whereas rat and mouse appear to have very similar substitution rates. Yet, the mouse and rat are very different biologically, despite their similar appearance. Consequently, the similarity combined with the different biology and a relatively long evolutionary distance provides unique comparative opportunities. Second, the rat is better characterized physiologically. Therefore, the opportunity to link the physiology of the rat to the mouse via comparative genomics is an advantage to the mouse community. Third, the ability to link the genetic power of the mouse to the rat via comparative genomics is a major help to the rat community. However, the most compelling argument is that the use of three mammalian species offers the opportunity to triangulate genetics, physiology, and clinical medicine. Sequencing the rat genome in parallel or immediately following the mouse provides the gold standard for comparative genomics and brings the nearly 200 years of pharmacological/toxicological data from the rat and drug discovery into genomic context with human and mouse.
Needs
Whereas the rat platform is steadily moving forward both in genetically defined strains and synthetic strains with transgenic technology, the rat system does require some additional infrastructure. At a recent rat priority meeting sponsored by Dr. Harold Varmus and the NHLBI, a small group of investigators were asked what additional infrastructure was needed for the rat. This 1-day meeting organized by the trans-NIH Rat Planning Group brought together >20 investigators from a broad range of disciplines. In addition, a survey was sent out to several thousand investigators to canvas the needs of the community. The results of this survey and the meeting participants generated the following partial list of priorities that need to be fulfilled:
- 1.
- Although the production of transgenic rats is routine, the creation of knockout has not been possible, as embryonic stem (ES) cells have not yet been possible in the rat. There is a critical need to develop ES cells or nuclear transfer (cloning) techniques in the rat.
- 2.
- There is a critical need to develop the Rat Genetic Resource Center to preserve and distribute the wealth of rat models to the research community.
- 3.
- Strengthening the Rat Model User Research Community via the Rat Community Forum Web site (http://taiji.ifrc.mcw.edu/RCF), rat-“specific” genomic/genetic-based meetings, and symposia on rats in conjunction with mouse and human genetics meetings is needed. Upcoming rat meetings are to be held at Cold Spring Harbor Laboratory, December 9–12 1999, followed by a meeting in Gothenberg, Sweden, July 2000.
- 4.
- Development of a third generation map consisting of single nucleotide polymorphisms (SNPs) would greatly facilitate the positional cloning of disease genes.
- 5.
- A BAC clone resource with an average insert size of 150 kb and 15-fold coverage will be needed for sequencing the rat genome.
- 6.
- The rat should be positioned for genomic sequencing as soon as possible.
Summary
Nearly 10 years ago, when I began building genomic tools for the rat as a postdoctoral fellow in the laboratory of Eric Lander, I was presented with a question: Which will occur faster, building the genomics for the rat or the physiology of the mouse? Now, I have a different view. Given the power of genomics and the strengths of various models, can we afford to exclude one model for another in advance of really knowing very much about gene function? It is critical to realize that no amount of ethylnitrosyl urea (ENU) or systematic knock out of all genes one at a time or in combination will replicate all of human biology in a single species. The baseline biology of any species provides strict, although, as of yet, undefined, boundaries that cannot be crossed. Consequently, the next millennium is likely to be dominated by groups that use the best models and systems to address specific questions. Research groups focused on a single species are likely to become extinct. Therefore, the increase in sequencing capacity and the continual drop in cost suggest that as many organisms as possible should be sequenced at least at the cDNA level, but scientists must present compelling arguments and a cohesive scientific community to lobby for the limited research dollars available for sequencing. As for the rat, it is a powerful platform for discovery of gene function. The basic tools are in place, and the resources required to “finish” the job are relatively minor compared with the investment already made in defining the biology and pathophysiology of this organism. The rat offers many advantages for identification of gene function that relate to common human diseases, because of the existing body of knowledge of physiological mechanisms, the availability of models that mimic these diseases, the ease of breeding, and the ability to generate new and better models that match subsets of patients at both the phenotypic and genomic levels (Gill et al. 1989;James and Lindpaintner 1997; Jacob 1999). Once genes and their functions are identified in rats, pathophysiologic mechanisms can be elucidated, and human genetic counterparts can be more easily identified.
Footnotes
-
E-MAIL Jacob{at}mcw.edu; FAX (414) 456-6516.
- Cold Spring Harbor Laboratory Press