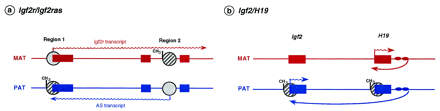
Expression competition mechanisms at the Igf2r/Igf2ras andIgf2/H19 loci. (a) Exons of Igf2r are depicted as solid boxes, with arrows indicating the transcribed alleles; regions 1 and 2 are sites of differential methylation (hatched circles with CH3: methylation) arising during embryonic development and in the egg, respectively. When the antisense transcript (AS) is expressed, the sense (Igf2r) is not and vice versa. The methylation mark at region 2 may function by regulating a putative antisense promoter that competes with Igf2r for expression (Barlow 1997); (b) H19 and Igf2 genes are indicated as solid boxes. The downstream H19endodermal-specific enhancers are represented by filled ovals. DMRs are represented by hatched circles; at the Igf2 gene the expressed allele is hypermethylated at two intronic regions (for simplification represented as a single region; there is also a maternally methylated region that is not shown here); methylation of the paternalH19 promoter blocks interaction with downstream enhancers on the paternal chromosome, which are then free to interact with the paternal Igf2 promoter (long arrow). Proper imprinting at theIgf2/H19 loci seems, therefore, to depend on promoter competition for the downstream H19 enhancers.











