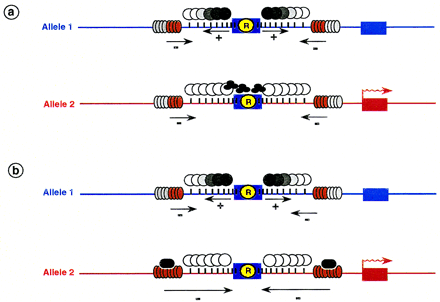
Hypothetical role for tandem repeats in the establishment of allele-specific methylation patterns at imprinted loci. These patterns are the result of the interaction of opposing positive (+) methylation and negative (−) demethylation signals. (a) The repeats (R) may act as methylation centers (see text), attracting or inducing the spread of methylation over a stretch of DNA (circles emanating from the repeat). The extent of methylation may be limited by counteracting demethylation signals (represented as gray/red disks) induced by CpG-rich environment (vertical bars). Trans-acting factors (trefoil structures) interfere with methylation or its spreading (allele 2) [a gradient of methylation is represented by differently shaded circles; (white circles) lack of methylation; (solid circles) complete methylation]. (b) Trans-acting factors (black ovals) may act alternatively in the demethylation pathway resulting in a dominant demethylation signal, spreading over the entire region (allele 2).











