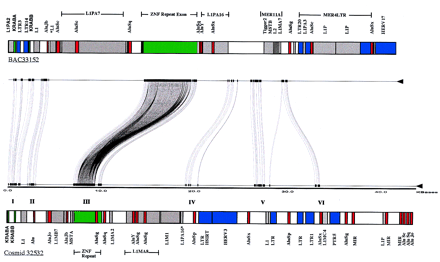
Comparative sequence analysis of ZNF duplicons. A genomic segment, corresponding to the entire insert (43,378 bp) of cosmid clone 32532 (GenBank accession no. HUAC003973) and paralogous positions 64,401–108,079 from BAC 33152 (GenBank accession no. HUAC004004), was compared between the two ZNF 19p12 clones using Miropeats software (threshold score s = 25; setting = onlyinter). Before analysis, common repeat elements were removed from the sequence using RepeatMasker software. Regions of sequence conservation that do not carry LINE or SINE sequences are delineated by “joining” lines between the two sequences. Six regions of paralogy were identified (indicated by Roman numerals), and each region was aligned using the BESTFIT alignment program (GCG software package). The position and identity of exons and other repeat elements are illustrated schematically above the miropeat alignment. Similar results were obtained using dot-matrix alignment software with unmasked sequence. Note that the identity and position of most repeat elements are not conserved between the duplicated segments.











