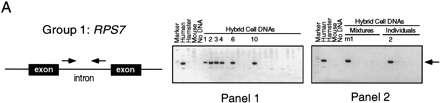
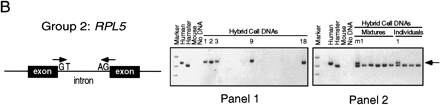
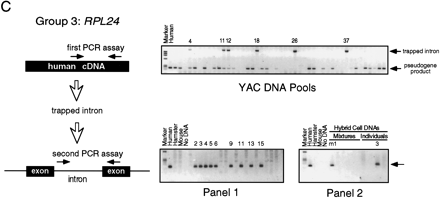
Intron trapping: derivation of STSs for and chromosomal assignment of representative group 1, group 2, and group 3 rp genes. (A) (Group 1) PCR primers were chosen from a previously sequenced intron of human RPS7. (Right) Results of testing of human/rodent somatic cell hybrid DNAs (NIGMS panels 1 and2) for the RPS7 STS; agarose gel stained with ethidium bromide. On each gel, the left-most lane contains size markers, and the next four lanes show results of PCR controls with human, hamster, or mouse genomic DNA, or no added DNA, as template. (Panel 1) 18 hybrid lines, each retaining multiple human chromosomes; 6 hybrids that tested positive for RPS7 are numbered; the results map RPS7 to chromosome 2. (Panel2) 24 hybrid lines, most retaining a single human chromosome; we initially tested these hybrids in pools of four (here mixture m1 is positive) and then tested individual hybrids from the positive pool (the chromosome 2 hybrid is positive). (B) (Group 2) PCR primers were chosen from human RPL5 cDNA sequence at a predicted splice site, with the dinucleotide GT appended to the 3′ end of the forward primer and the dinucleotide CT (the complement of AG) appended to the 3′ end of the reverse primer. (Right) Results of hybrid mapping, which assigned RPL5 to chromosome 1. The arrowhead (extreme right) indicates the size of the human PCR product; a smaller, hamster product is also present in many lanes. (C) (Group 3) Mapping of RPL24 involved two quite different PCR assays. The first PCR assay (top), with 45 YAC DNA pools as template, was designed to trap an intron with primers chosen from human cDNA sequence according to rules discussed in the text. Six YAC pools that yielded the higher molecular weight, trapped-intron product are numbered; many more pools yielded the lower molecular weight, pseudogene product. The control reaction with human genomic DNA as template yelded only the pseudogene product. Sequencing of the trapped intron made possible a second PCR assay specific to the functional RPL24 gene (bottom); with the second assay, the gene was mapped to chromosome 3.











