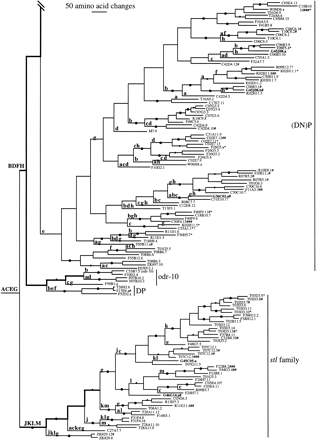
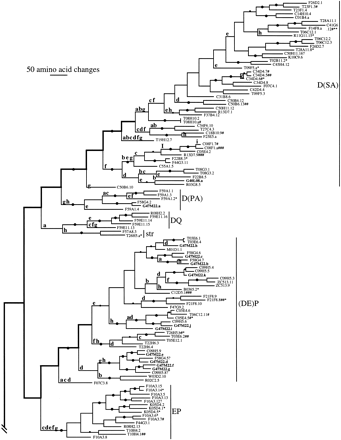
Phylogenetic tree relating 257 members of the str andstl families of chemoreceptors. The str subfamilies are indicated. Branch lengths are proportional to the number of inferred amino acid changes. Bootstrap support above 75% is indicated by a large dot on the branch supporting the relevant node, with a small dot indicating bootstrap support of 50%–75%. Lowercase letters above branches indicate inferred intron loss, whereas uppercase letters indicate intron gain. Double thickness lines connect hypothetical ancestral genes inferred to have retained the full complement of eight introns in each family. C. briggsae genes are indicated by boldface type and all start with the letter G. Pseudogene status is indicated by symbols after each gene name: (?) Loss of start codon or questionable intron boundary; (*) in-frame stop codon; (#) frameshift or large indel.











