WebWise: Guide to the Baylor College of Medicine Human Genome Sequencing Center’s Web Site
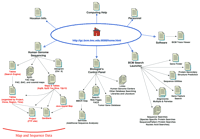
This installment of the WebWise series reviews the Baylor College of Medicine Human Genome Sequencing Center’s web site. When comparing this web site to the site map depicted in Figure1, you may notice that some duplicate and minor links have been omitted. The main features of this web site, as well as the features of those sites already reviewed, are indicated in Table1.
The Baylor College of Medicine Human Genome Sequencing Center Site Map. The main links to pages discussed in the text are illustrated here. Links on the top of the Home Page are to general informational resources; links located on the bottom portion are to the data and additional tools.
Features of the BCM Web Site
General Information
The Baylor College of Medicine Human Genome Sequencing Center (BCM) Home page (http://gc.bcm.tmc.edu:8088/home.html) presents a very straightforward organization. There are four stylized buttons linking to sequence data, analysis, and tools pages followed by stylized buttons linking to more general information such as the Genome Sequencing Center Personnel, BCM Departments, Computer Help, and Information about Houston (including maps). The Genome Center Computer Help page appears to be primarily for internal BCM use as it includes some information targeted toward people having a BCM account. The Genome Sequencing Center Personnel page is useful if you need contact information (http://kiwi.imgen.bcm.tmc.edu:8088/personnel.html). This page lists the names, titles, e-mail addresses, and phone numbers of the Center’s personnel from the Director, Dr. Richard Gibbs, on down. The Usage Statistics link located at the bottom of the Home page brings you to a page that should have current usage statistics but has not been updated in a long time.
Data
Follow the BCM Human Genome Sequencing link from the Home page to access the sequence data. The BCM sequencing effort is focused on two regions of the X chromosome (Xq28 and a ∼35-Mb region of Xp22) and one region of chromosome 12 (12p13). A general overview of the community-wide sequencing effort is available for the X chromosome by following the Ideogram link. This is a comprehensive overview of the overall sequencing effort for the X chromosome. Unfortunately, some of the text is too small, making it difficult to read, and a similar overview is not provided for the chromosome 12 target. It would be useful to have an indication of how current the data in the ideogram are and a more explicit statement on the total amount (in Mb) of human DNA sequence data already produced by, and projected to be produced by, the BCM. Admittedly, this type of information is not of paramount importance, but it is still a useful summary, when available, and does serve to satisfy a certain curiosity about the overall size of the sequencing effort undertaken by any given center.
BCM has made the effort to integrate the map and sequence data by presenting both physical maps and tables for all three targeted regions (Xq28, Xp22, and 12p13). Integrating map and sequence data is a prerequisite to obtaining a full comprehension of the sequence progress, and unfortunately some centers do not provide a graphic view of the sequenced clones. The maps displayed on these web pages all include the critical information—the location, name, and status of the sequencing clones; however, they use different map styles for different targets, which requires a greater effort on the part of the viewer to interpret. Furthermore, the majority of these map displays are significantly wider than the average browser window, which forces one to use both vertical and horizontal scrolling to see the whole display.
There are 10 data-relevant links available on the Human Genome Sequencing page (http://gc.bcm.tmc.edu:8088/cgi-bin/seq/home); some of these links lead to pages containing a table of sequencing results, a static physical map, and/or a link to an interactive Java map view (the links to Xq28, 12p13, and the four Xp22 YAC-defined bins; Maps & Tables in Fig. 1). The 12p13 page does not include a physical map display directly on the page but, instead, relies on the Java Applet map display, which does not include all of the data provided in the table. The Tables link, located toward the bottom of the Human Genome Sequencing page, leads to a search engine that will build a table of data for individual clones, projects, and regional targets or list data for the entire BCM sequencing effort (http://gc.bcm.tmc.edu:8088/cgi-bin/seq/bcm-web-tables). This search tool works well, provides flexible options, and should provide a useful method to track sequencing progress once a clone, project, or regional target name is identified from the maps or data tables. The remaining links (YAC map, and PAC, BAC, and cosmid maps) lead to more detailed physical map views without the corresponding tables of sequencing results. When one first visits the Human Genome Sequencing page, it is not immediately obvious which links lead to DNA sequencing results. Although, in general, brevity is a plus on the web, a minimal description and/or more explicit link nomenclature can facilitate navigation.
Although there is a direct correspondence between the maps and tables, it is not fully intuitive because (1) the tables of data are initially confusing, and (2) three different map styles are used. Before extracting information from the tables or physical map displays, one must first take a moment to become familiar with the format and nomenclature. On the plus side, the tables that are associated directly with a map display and the tables generated with the search tool do utilize a uniform format. The eight columns in the table provide the project name, GenBank accession number (when available), clone name(s), vector, region (e.g., Xp28), description, size, and status. DNA sequence data are accessed by following the hot-linked GenBank accession number to the NCBI GenBank record or by following the linked Project Name to the sequence on BCM’s FTP site. Additional hot links provided in the GenBank, Description, and Status columns all lead to a table generated by the search tool. For example, project J-15 (Xq28 region) refers one to project J-28 in the GenBank and Status columns. These two J-28 links, as well as the linked Description text, all call up the table for project J-28 that is generated by the search tool (follow the Table link from the Genome Sequencing page). Each clone being sequenced is assigned a project name. As clones are put into contigs, they are reidentified as a Joined project and assigned a new project name (Joined project names are prefixed with a J). As the projects are further joined together (e.g., as larger contigs are formed), the process is repeated and a new project name is assigned. The tables give a misleading first impression of the quantity of sequence data available as they list all of the various projects a clone has progressed through, instead of just defining the final contig. Presenting all of the available data in this manner allows one to reconstruct the process used to build contigs, but also results in a more cumbersome display that includes a considerable amount of redundant data.
Three different styles are used to depict the physical maps. The map available for Xq28 uses one style, the maps available for the Xp22 region utilize a second style, and all three regions (Xp22, Xq28, and 12p13) include a link to a third map view, namely, an experimental Java-based interactive map display that is not yet ready for full-scale use. For example, the Java-based map still has the following limitations: the Show Sequence Map button does not appear to function, there is not full correspondence between the clones listed in the table and those appearing on the Java map display, and some project names are not indicated (e.g., the Joined projects). On the plus side, the display itself is a pleasingly simple color-coded representation of the clone tiling path and sequence status. In addition, one can manipulate the scale of the map (e.g., zoom in/out), which facilitates browsing at both the general overview and detail-oriented levels.
The two static map displays for the Xq28 and Xp22 physical maps use color-coded rectanges or lines, respectively, to denote sequence status and indicate both the clone and project names. On the Xq28 map, clone names are indicated below each rectangle, and the original project name is shown in boldface type above the rectangle. Unfortunately, the Joined project names are not explicitly indicated on this map, but this information can be deduced by examining the table. In contrast, the Xp22 maps do indicate the Joined projects. The color key is not included directly on the Xp22 diagrams but, presumably, is the same key as the one available for the PAC, BAC, and cosmid physical maps, as these maps use the same format. Presenting three different map formats, each of which uses a different color key, can make it more challenging to use these maps; however, by examining both the tables and maps one can determine the clone order and sequence status for each region being sequenced.
Tools
With regard to tools, the BCM web site is an interesting study in contrasts. A host of special features is offered, as outlined below, yet there is minimal attention paid to describing the software and/or automation tools being used in-house for genomic DNA sequence generation and analysis. And, although they have made available a very useful sequence data search tool (Tables; see above), they do not provide any capability to carry out a homology comparison against the DNA sequence data they have generated. This is rather remarkable, given the collection of analysis tools offered via the BCM Search Launcher and the Biologist’s Control Panel (see below). Admittedly, this is perhaps a contentious issue. It does require a certain level of expertise to set up and maintain a local BLAST server, and the majority of their sequence data are already available in GenBank and thus available for comparison via NCBI’s BLAST servers. Yet, people continue to perceive the sequencing center’s web sites as the best source of the most up-to-date sequence data, and given that opinion, homology searching of the local data becomes an important feature.
One software tool being developed at the BCM is described on the Software page (follow the Software link from the Home page;http://gc.bcm.tmc.edu:8088/software/software.html) and will be ready for distribution soon. A nice Java demonstration of the BCM Trace Viewer is embedded in the web page—try clicking on the buttons and moving the scroll bars around. As the name implies, the Trace Viewer displays the trace, base calls, and quality values. This tool seems to afford a lot of flexibility; it can be run as a stand-alone application, or embedded in a web page, and the user has a lot of control over the display.
Although the immediate goal of the Human Genome Project is DNA sequence generation, the impact of these data is an expanding need for sequence analysis strategies and tools. Increasingly, web sites hosted by sequencing centers and other “genomics” groups are including some sequence search and/or analysis tools. Although this is a wonderful service to provide, free, to the research community, the size and complexity of the World Wide Web can make it challenging to locate these resources. The BCM has obviously recognized both the importance of making search and analysis tools available via the web, and the difficulties inherent in finding disparate resources on the web. To address this issue they have made a significant commitment to provide an integrated set of molecular biology sequence search and analysis tools on their web site.
The BCM Search Launcher (Smith et al. 1996;http://gc.bcm.tmc.edu:8088/search-launcher/launcher.html) provides a single starting point via a convenient web interface for a large variety of sequence search and analysis tools. Many of these tools are also available elsewhere, either at the location where they were originally developed or at a secondary host site. The main advantage to the BCM Search Launcher is that it presents these tools as a single collection at one site, thus obviating the need for extensive web surfing to first locate and then utilize these tools. The main disadvantage is that the options are not fully integrated; one must try each tool successively, and become knowledgeable about each different resource used. These tools fall into four general categories: (1) protein analysis; (2) nucleic acid analysis; (3) sequence manipulation tools; and (4) sequence alignment. The BCM Search Launcher consists of a collection of several different tools for a given type of analysis, where each of these collections is called a Launch Page. Each Launch Page adopts the same general format, where a text box into which you paste your sequence is located toward the top of the page and the different available search options are listed under that. One must simply paste in a sequence, check off the Radio button for the desired option, click on the Submit button, and await the result (results are returned via the web interface for the majority of the analysis tools available here).
Some of the BCM Search Launcher options for protein analysis include homology searching (including some species-specific searches), pattern finding, alignments (pairwise or multiple), and secondary structure prediction. The nucleic acid analysis options include homology searches, gene feature searches, and pairwise or multiple sequence alignments. Several options are available on the Gene Feature Launch page, and one additional tool, the BCM Gene Finder, is also available on the BCM web site but is not integrated into the Launch page. The Gene Finder can be accessed either via a link on the BCM Search Launcher page or via the Biologist Control Panel page (a link to this is provided on the Home page). The Sequence Utilities Search Launcher page includes various sequence manipulation tools such as converting a sequence to FASTA format, PCR primer prediction, restriction mapping a sequence, translating the sequence, or generating the reverse complement of the sequence. Although the functionality of these tools was not tested for the purpose of this review, this is an impressive collection of DNA and protein search and analysis tools.
Some additional resources are listed on the Biologist’s Control Panel page (http://gc.bcm.tmc.edu:8088/bio/bio_home.html). Much of this page consists of an extensive collection of hot links to other web sites, but one other interesting resource is available by following the MBCR Help link near the bottom of the page. From the MBCR page (Molecular BiologyComputational Resources) follow the Databanks or Services link; these two pages list numerous additional sequence analysis tools that are available on the web but are not (yet) included in the BCM Search Launcher.
Conclusions
In addition to the human DNA sequence data generated by this group, the BCM Human Genome Sequencing web site provides a considerable collection of sequence analysis resources to the research community. Assuming that all of these tools work as expected (to reiterate, function was not assessed), this resource provides a starting point for exploratory analysis of sequence data. Given the emphasis on tools and resources evidenced by the BCM Search Launcher and the Biologist’s Control Panel, it is surprising to note the absence of both (1) the capability to carry out homology comparison searches against the BCM DNA sequence data, and (2) documentation of the software and/or automation tools utilized to generate and manage the DNA sequence data. Data management is a significant factor in the Human Genome Project, and as sequence production is scaled up over the next several years, the importance of this factor will only increase.
As a whole, the organization, style, and link nomenclature used on this web site is somewhat uneven. The Home page is organized well, and the link nomenclature used is meaningful, making it easy to determine what the general content is behind each link. However, the links available on the Human Genome Sequencing pages do not indicate content as explicitly, so a first-time visitor might end up browsing through several of the Xp22 links before finding the table of sequencing results. The majority of the pages include a link at the bottom of the page back to the Home page, but a consistent set of internal navigation links is not available here. On those pages where some internal navigation links are provided, for example, the Biologist’s Control Panel, some of the links are either broken or obsolete (e.g., the Newsletter and GRM links). While acknowledging that web page style does generate diverse personal opinions, in general, this web site uses the style format of “first-generation” web sites (Siegel 1997). For example, the page background is gray, and heavy borders are used on all of the tables. This web site is rich in content, but a few simple changes would enhance the presentation, which in turn would facilitate access.
Commendably, sequence data are integrated with map data. However, the oversized map format does hamper information retrieval. Apparently the BCM is aware of some of these limitations as they continue to explore the use of Java applets to depict map data. All of the physical map diagrams depict the clone name, order, and status of sequenced clones—critical information needed to form a “big picture” of the sequence status in a region of interest. Without maps of this nature, it can be exceedingly difficult to determine the clone tiling path, which is essential information for anyone trying to reconstruct the DNA sequence in a region harboring a gene of interest.
Footnotes
-
↵1 E-MAIL ; FAX (301) 435-2433.
-
Next Month: The Genome Sequencing Centre Jena
- Cold Spring Harbor Laboratory Press