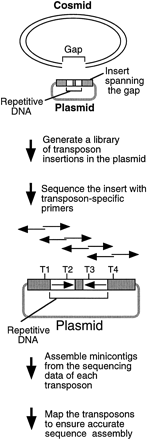
Strategy for sequencing repetitive DNA. A plasmid is identified with an insert spanning the cosmid gap, a transposon library is generated, and the insert is sequenced using the transposons. T1–T4 represent transposon insertions. The two sequencing reactions initiated from individual transposon insertions are shown as back-to-back arrows, which can be assembled using the 5-bp target site duplications. The transposon order can be determined by mapping to ensure accurate sequence assembly.











