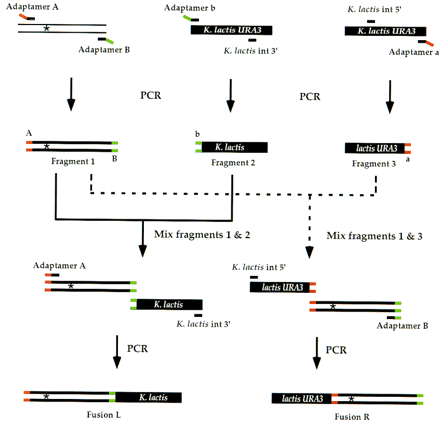
Generating fusion fragments for allele replacement. The gene of interest with an altered site [indicated by an asterisk (*)] is amplified by PCR using adaptamers A and B (fragment 1). Similarly, two overlapping K. lactis URA3 fragments are generated separately by PCR with the K. lactis URA3 adaptamers and two internal K. lactis URA3 primers (fragments 2 and 3). Fragments 2 and 3 do not encode full-length URA3 and thus are represented as K. lactis and lactis URA3,respectively. The ends tagged by adaptamers A, B, a, andb are labeled on the initial PCR products. As described in Fig.1, fragments 1 and 2 are mixed with the K. lactis int 3′ primer and adaptamer A for an additional PCR step to generate a new fusion product (fusion L). In a separate PCR, fragments 1 and 3 are mixed with the K. lactis int 5′ primer and adaptamerB to generate a second chimeric fragment (fusion R). Both fusions L and R contain the altered site.











