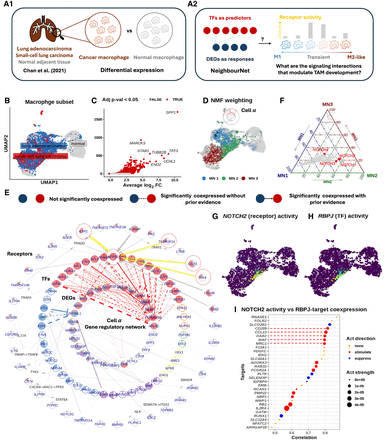
Prior knowledge annotation highlights critical signaling interactions facilitate tumor-associated macrophage development. (A) (A1) We focused on the macrophage subset of Chan et al. (2021)’s small cell lung cancer scRNA-seq data set. Using NNet, we aimed to identify the signaling interactions that regulate differentially expressed genes (DEGs) between cancer and normal macrophages, providing insights into how inter- and intracellular signaling interactions shape macrophage identity within the tumor microenvironment. (A2) We utilized NNet’s upstream signaling pathway (USP) inference to identify receptors with high activity in driving TF regulation of DEGs in transient macrophage populations during TAM development. (B) UMAP visualization of macrophages in the data set. (C) Volcano plot of the DE result. Only genes that were upregulated in cancer are shown. Pi value: negative log10 p-value times log2 fold change. (D) Soft clusters of cells for the first three meta-networks, illustrating a key transition from proinflammatory macrophages to TAMs. Cell α, with the highest cell weight on meta-network 1, is highlighted. (E) Annotated coexpression network of cell α. The innermost layer contains response (target) genes, encircled by central TFs in the meta-networks. Receptors occupy the outermost layer, each linked to a TF inferred to mediate that receptor’s influence. If the receptor–TF link is indirect, an extra layer shows the shortest signaling path according to prior knowledge. The Methods section provides details. This network captures the canonical NF-kB pathway in proinflammatory macrophages. (F) Ternary plot of receptor activity scores on the first three meta-networks. NOTCH receptors (NOTCH2/3/4) show peak activity in meta-network 2. (G) NOTCH2 activity and (H) RBPJ connectivity projected onto the UMAP. RBPJ was the TF whose connectivity was found mostly correlated with NOTCH2 activity. (I) Correlation between NOTCH2 activity and different targets’ coexpression with RBPJ. The top targets with the highest correlations are illustrated. NOTCH2 activity is strongly associated with M2-like TAM marker expression mediated by RBPJ.











