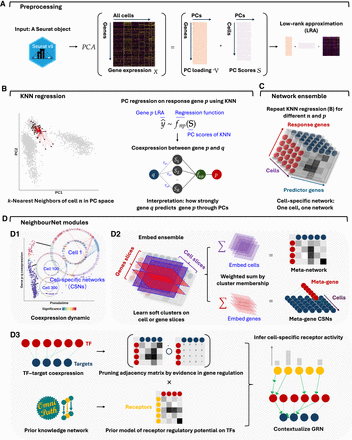
NNet workflow for inferring cell-specific coexpression networks and integrating prior knowledge. (A) Preprocessing: Single-cell gene expression data are subject to PCA to capture the major variation. A low-rank approximation (LRA) then reconstructs the expression matrix using the PCs. A weighted k-nearest neighbor (KNN) graph that defines each cell’s local neighborhood in the PC space is constructed. (B) Neighborhood regression: For each cell, NNet performs regression within the cell’s KNN using the LRA-derived gene expression as the response and PCs as predictors. This process quantifies coexpression between genes by measuring how “predictor genes” (those used to embed PCs) contribute to predict each response in the PC space. Repeating this for multiple response genes yields cell-specific coexpression networks (CSNs). (C) Network ensemble construction: Collecting all CSNs form a network ensemble, which is a cell × response × predictor 3D array that can be diced and sliced. (D) Downstream analysis: (D1) We can slice the network ensemble to examine CSNs, analyzing cellular variation and gene regulation dynamics through coexpression. (D2) Nonnegative matrix factorization (NMF) identifies soft clusters of cells with similar network structures. Aggregating CSNs by these clusters yields meta-networks that summarize shared coexpression patterns. Same analysis strategy can be applied to the gene dimension corresponding to TFs, producing TF modules that coregulate common programs. Aggregating TFs by module yields simplified meta-TF–target networks with improved interpretability. (D3) We adapted the NicheNet framework to integrate gene regulation and signaling interaction databases from OmniPath, constructing integrated prior knowledge networks (PKNs). Annotating CSNs with these PKNs transforms them into contextualized gene regulatory networks (GRNs). In addition, PKNs enable the inference of upstream signaling pathways (USPs) for each CSN by tracing signal transduction paths (receptor–TF–target) inferred based on the contextualized TF–target interactions.











