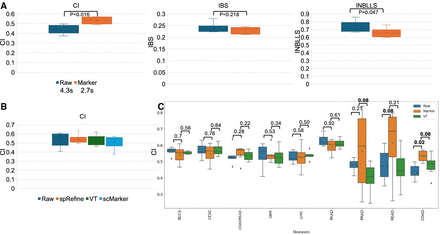
Applications of spRefine for disease modeling. (A) Comparison of survival prediction performances based on three different metrics between raw expression profiles and expression profiles only containing genes selected by spRefine. The cancer type of selected data is COAD. We highlighted the running time comparison and the Wilcoxon rank-sum test (one-side) p-value (P) in this panel. (B) Comparison of survival prediction performances across marker genes from different sources. (C) Comparison of survival prediction performances across different gene sets based on the cancer types from different cancer types. We annotated the Wilcoxon rank-sum test (one-side) p-value (P) in this panel, and highlighted the significant result (p − value < 0.1).











