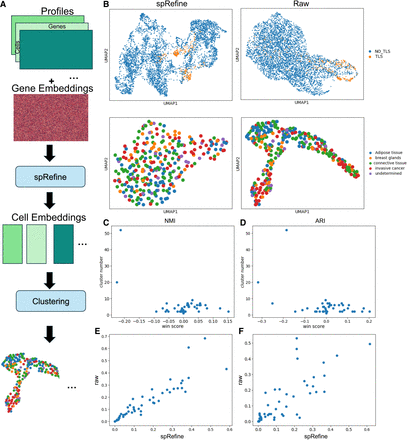
Analyzing the results of spRefine pretrained with large-scale spatial transcriptomics. (A) The workflow of pretraining spRefine for phenotype-level identification. (B) The UMAPs for visually comparing spot representations before (right) and after (left) imputation based on the two selected samples with different cell states. (C) The relationship between win rate computed based on NMI and cluster number. (D) The relationship between win rate computed based on ARI and cluster number. (E) The relationship between the NMI scores of raw data and data imputed by spRefine. (F) The relationship between the ARI scores of raw data and data imputed by spRefine.











