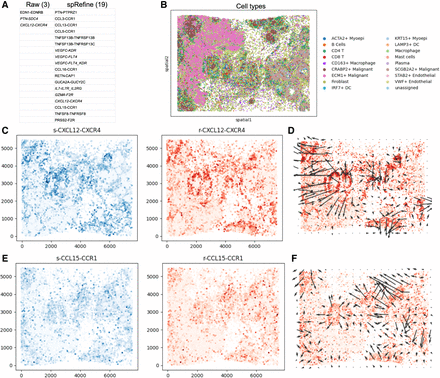
Using spRefine for discovering novel cell–cell communications (CCCs). (A) The comparison of discovered signals between the results of COMMOT based on the raw expression profiles (Raw) and imputed expression profiles (spRefine). In (C) and (E), “s” represents sender and “r” represents receiver. (B) The visualization of cells in xenium_breast data set colored by cell types based on spatial location. (C) The plots containing sender and receiver strength for the CCC CXCL12-CXCR4. (D) The signal direction of the CCC CXCL12-CXCR4. (E) The plots containing sender and receiver strength for the CCC CCL15-CCR1. (F) The signal direction of the CCC CCL15-CCR1.











