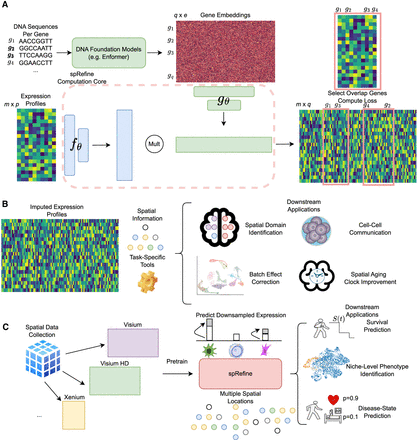
The landscape of spRefine and related applications. (A) The computation framework of spRefine as a tool for imputation and denoising. spRefine utilizes prior information from DNA foundation models to help uncover missing gene–gene interactions and gene expressions unmeasured by the raw transcriptomic profiles. (B) The applications of spRefine including spatial domain identification, CCC inference, batch effect correction, and spatial aging clock improvement. (C) spRefine as a pretraining framework for representing phenotype information. spRefine is capable of performing survival prediction, identifying spot-level phenotype information, and predicting disease states.











