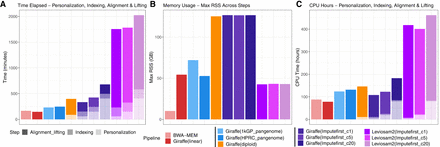
Computational efficiency of personalization, indexing, alignment, and lifting across the evaluated workflows. (A–C) Summarization of wall-clock runtime, peak memory usage, and CPU time for linear (BWA-MEM, Giraffe(linear)), pangenome (Giraffe(1 kGP_pangenome), Giraffe(HPRC_pangenome)), personalized-pangenome (Giraffe(diploid)), and impute-first strategies using either VG Giraffe (Giraffe(Imputefirst_c5), Giraffe(Imputefirst_c20)) or LevioSAM2 (Leviosam2(Imputefirst_c5), Leviosam2(Imputefirst_c20)). Bars are shaded to distinguish the types of computation. Note that only the workflow steps that depend on the input data are measured here. “Offline” index building steps that are input data independent are not counted. All workflows were executed using 32 threads, except for the BCFtools consensus and BWA-MEM indexing steps, which ran on a single thread.











