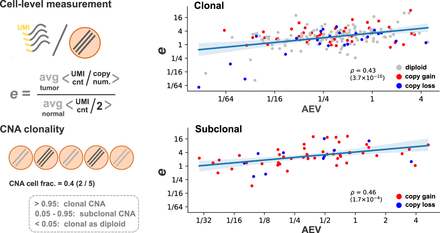
Single-cell model evaluation. Five cutaneous squamous cell carcinoma samples with scRNA-seq and WES were reanalyzed based on the cell type annotation by Ji et al. (2020). In each sample, unique molecular identifier (UMI) counts per gene copy were averaged over cell populations annotated as tumor or normal. The ratio of the tumor average to normal was used as the copy-number-adjusted expression difference between tumor and normal tissues (the e parameter). The clonality of CNA events was defined by a fraction of tumor cells carrying the alteration. For substitutions identified from WES, AEV and e values were plotted separately for 198 cases in which the background ploidy is clonal and 61 subclonal counterparts (median CNA cell fraction: 0.31).











