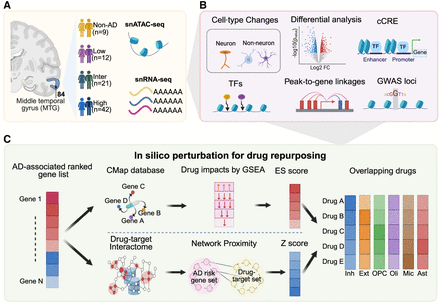
Overview of study design. (A) Collection and preprocessing of snRNA-seq and snATAC-seq data sets in physical space. (B) Multiple analysis to identify an AD-associated gene list including differential gene expression analysis; identification of cCRE; cell type–specific, AD-associated risk variants; and peak-to-gene linkages. (C) Two in silico perturbation methods for drug repurposing. Network proximity analysis identifies potential drugs by assessing the proximity between the drug target set and AD-related gene set. Drug efficacy is evaluated based on the proximity distance and Z-score. GSEA leverages drug–gene signatures from the CMap database and differentially expressed genes to calculate the enrichment score (ES). The ES reflects the drug's potential to reverse the observed gene expression patterns within the given network.











