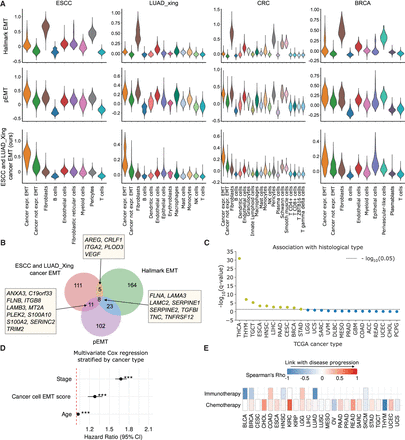
Application of ANS to devise EMT signature specific to malignant cells. (A) Score distributions of the Hallmark EMT, the pEMT, and our proposed ESCC-specific and LUAD_Xing-specific cancer EMT gene signatures in different cell types of ESCC, LUAD_Xing, CRC, and BRCA. (B) Venn diagram for the overlap in gene lists for the Hallmark EMT, pEMT signatures, and the signature we designed. (C) Association of ESCC-specific and LUAD_Xing-specific cancer EMT signature scores and histological subtypes in TCGA (only cancer types with at least one histotype are included). Significant associations (Q value < 0.05) are represented by dots above the dotted line (yellow). The FDR-adjusted P-values from the two-sided Kruskal–Wallis test were −log10-transformed and sorted in decreasing value, with a higher y-value indicating higher significance. (D) Multivariate Cox survival analysis with disease stage, patient age, and cancer cell–specific EMT signature scores calculated in bulk TCGA RNA-seq data. The model was stratified by cancer type. Variable-specific P-value significance, (***) P-value < 0.001, (**) P-value < 0.01, (*) P-value < 0.05. (E) Association between the cancer cell–specific EMT scores in bulk TCGA tumors and patient treatment response. Association was calculated with Spearman's rank correlation between the scores and treatment response ranked one (complete response) to four (clinical progressive disease; Methods).











