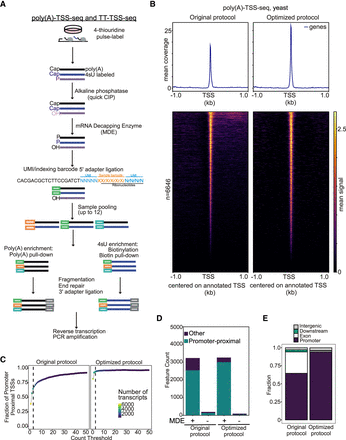
Optimization of the TSS-seq protocol with yeast RNA. (A) Schematic of the TSS-sequencing protocol. Alkaline phosphatase (quickCIP) removes the 5′-phosphate groups from noncapped fragments (shown in purple). The mRNA decapping enzyme (MDE) removes the 5′-terminal caps, exposing a 5′-phosphate group (labeled P). This enables the ligation of a 5′ adaptor, which contains sample barcodes to allow sample multiplexing and UMIs. Nascent (4 thiouridine (4sU), labeled blue) or steady-state RNA (poly(A), labeled black) is then selected, before fragmentation, end-repair, 3′ adaptor ligation, reverse transcription, and PCR amplification. (B) Density plot and heat map showing tag locations compared to annotated TSSs for poly(A)-TSS-seq. RNA isolated from yeast was subjected to poly(A)-TSS-seq using either the original or optimized conditions. The x-axis is centered on the annotated TSS. (C) Fraction (y-axis) of tags located in promoter-proximal regions (–300 to +100 bp from annotated ORF start codons) at different read thresholds (x-axis). (D) Bar plot showing the number of tags located in promoter-proximal regions with the original or optimized poly(A)-TSS-seq protocols (+MDE). As negative controls, samples that had not undergone the decapping reaction were included (−MDE). A threshold of n = 3 counts was applied. (E) Genomic locations of detected tags. Tag counts were normalized using the DESeq2 median-of-ratios approach using a threshold of n = 3 counts.











