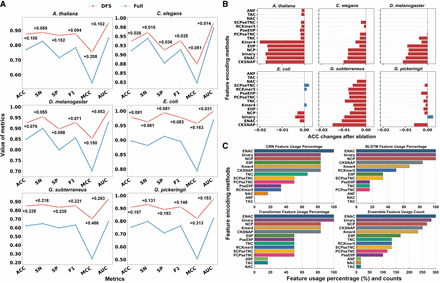
Figure 2.
The impact of feature selection and encoding on EnDeep4mC's performance. (A) Metrics of the ensemble model using DFS versus full 14 encodings on the data sets of six species. (B) ACC changes of features selected by the DFS framework after ablation (Supplemental Tables S8–S10). (C) The selection rates of 14 feature encoding schemes on different base models and the total usage frequencies on ensemble model.











