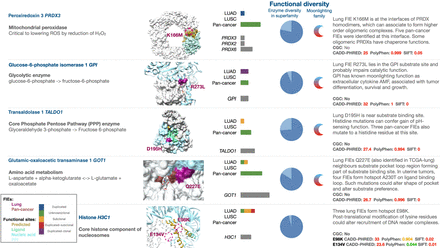
FIE genes in functionally diverse protein families are targets for discovery of potential neofunctional events late in tumor evolution. Selected examples show postduplication or subclonal lung FIEs in metabolic enzymes from functionally diverse CATH superfamilies (PRDX3, GPI, GOT1, and TALDO1) and an oncohistone (H3C1). Functional diversity: Enzyme diversity in superfamily indicates the number of distinct enzyme functions in the FIE's superfamily as a fraction (light blue) of the most diverse superfamily. Moonlighting family indicates when at least one gene has previously identified moonlighting functions. Bar plots indicate the distribution of FIEs identified in LUAD, LUSC, and pancancer, color-coded by the presence of gene duplication and mutation clonality as per legend. Pathogenicity predictions are shown in red (above the respective pathogenicity threshold for CADD, PolyPhen, or SIFT), in orange if they are possibly pathogenic, and in green otherwise. Oncohistones were identified by manual curation of nonmetabolic genes. Larger-sized structures are shown in Supplemental Figure 7. A complete list of 28 diverse functional families is provided in Supplemental Table 5.











