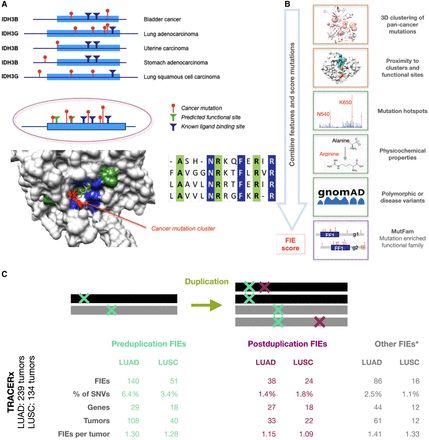
FunVar protocol identifies FIEs in lung tumor evolution. (A) FunVar protocol. Mutations (red) from different tumor types and gene paralogs were grouped when they shared protein domains predicted to have equivalent functions because they belong to the same functional family in the CATH database (Sillitoe et al. 2015; Das et al. 2015b). These mutations can be mapped to a single 3D structural representative. Known functional sites from paralogs (blue), and likely functional sites predicted owing to high conservation within the CATH functional families (green), are mapped in an identical manner. Significant mutation clusters near functional sites were termed “tunable sites,” highlighting commonalities between mutations from different cancer types and paralogs via their impacts on specific protein functions and aiding the detection of rare events. (B) FunVar scoring. TRACERx lung mutations were tested for occurrence in tunable sites and given functional impact event (FIE) scores using a simple heuristic based on mutation properties. (C) FIEs identified pre- and postduplication for LUAD and LUSC. Number of FIEs, FIEs as percentage of SNVs (missense and synonymous), number of distinct genes containing FIEs; number of tumors with FIEs, and average FIEs per tumor. Other FIEs occur in regions with no gene duplication, although a minority of these occur in areas of monoallelic duplication when the timing of the mutation relative to the duplication may be unknown.











