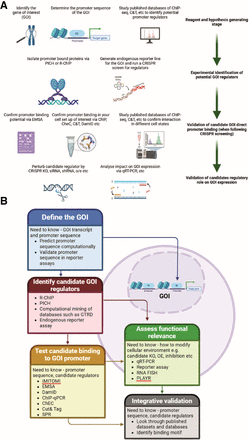
Integrated workflow for identifying direct promoter-bound regulators of a GOI. (A) This schematic outlines a stepwise strategy for decoding transcriptional regulation of a GOI, with a focus on identifying direct regulators acting at the promoter level. The process begins by selecting the GOI, including annotation of transcript variants, promoter structure, and determining regulatory elements. At this stage, mining publicly available data sets, such as ChIP-seq, CUT&Tag, and DamID profiles, can yield initial hypotheses regarding candidate transcription factors or chromatin regulators. Next, unbiased identification of promoter-associated proteins can be performed using locus-specific proteomics techniques such as PiCh or reverse ChIP (r-ChIP). In parallel, genome-wide CRISPR knockout screens coupled to endogenous reporters (e.g., fluorescent tagging or PLAYR-based detection) can be employed to identify both direct and indirect regulators of GOI expression. Following candidate identification, direct promoter occupancy should be validated using complementary in vitro and in vivo binding assays, including electrophoretic mobility shift assay (EMSA), ChIP, cleavage under targets and release using nuclease (CUT&RUN or CHEC), and DNA adenine methyltransferase identification (DamID), complemented by systematic evaluation of available public data sets. This step ensures that the regulators physically interact with the GOI promoter and are not acting solely via indirect pathways; consequently, it is critical to perform this analysis if candidate identification occurred via CRISPR screen as this will also identify non-promoter-bound indirect regulators. Finally, functional validation needs to be performed by perturbing candidate regulator levels—via overexpression, RNAi, or CRISPR-based approaches—and quantifying changes in GOI expression using qRT-PCR, RNA-FISH, or related transcript-level assays. Importantly, all assays should be conducted in physiologically relevant cellular contexts to preserve native chromatin architecture and regulatory interactions. (B) This iterative framework integrates discovery, validation, and functional characterization of promoter-bound regulatory factors and can be adapted to dissect transcriptional control of diverse genomic loci.











