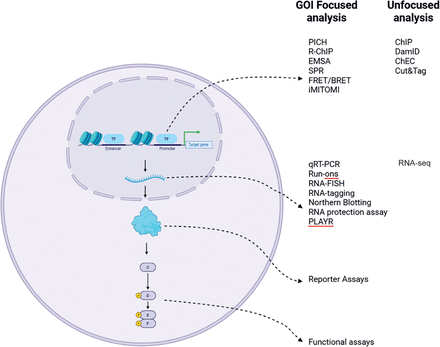
Molecular assays used to investigate gene regulatory mechanisms, broadly categorized into four functional levels: (1) transcription factor (TF)–DNA binding, (2) mRNA abundance of the GOI, (3) GOI protein expression, and (4) GOI-associated signaling networks. (1) TF–DNA binding: Assays at this level, such as ChIP-seq, CUT&RUN, or EMSAs, provide direct evidence of a physical interaction between a TF and the regulatory elements of the GOI. These approaches assess binding specificity and occupancy but do not inform on the functional consequences of such binding. (2) GOI mRNA levels: Quantification of GOI transcripts (e.g., via RT-qPCR or RNA-seq) offers a measure of transcriptional output. Changes in mRNA abundance in response to TF perturbation can suggest regulatory influence but may still reflect indirect effects mediated by intermediate factors. (3) GOI protein levels: Techniques such as reporter assay or western blotting/mass spectrometry–based proteomics assess the steady-state levels of the GOI protein. Although changes at this level may reflect transcriptional regulation, they are also influenced by post-transcriptional and translational mechanisms, including mRNA stability, translation efficiency, and protein degradation rates. (4) GOI signaling networks: This includes analysis of downstream signaling pathways or phenotypic outputs modulated by the GOI protein. These readouts are inherently multilayered and integrate regulatory events occurring at several preceding molecular levels. It is critical to recognize that as we move downstream in the gene expression cascade, from TF binding to phenotypic outcome, additional layers of regulation become increasingly prominent. For instance, using GOI protein levels as a surrogate for transcriptional regulation by TF X can be misleading. An observed increase in GOI protein levels following TF X overexpression does not necessarily imply direct TF–GOI transcriptional activation. Instead, TF X may indirectly enhance GOI protein abundance by modulating expression of genes that encode mRNA-stabilizing factors, translation enhancers, or protein chaperones that extend GOI half-life. Thus, careful interpretation of molecular readouts is essential, and conclusions regarding direct regulatory relationships should be supported by complementary mechanistic evidence across multiple levels of analysis.











