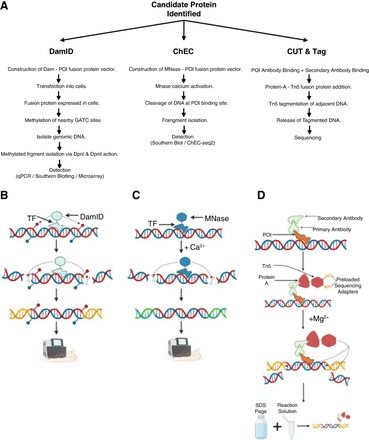
Comparison between select protein focused protein–DNA interaction analysis methods. (A) DamID (Greil et al. 2006; Aughey and Southall 2016), ChEC (Schmid et al. 2004), and CUT & Tag (Kaya-Okur et al. 2019) workflows following candidate protein identification. (B) DamID-sequencing. DamID-fusion with candidate TF binds to DNA and proceeds to methylate nearby adenines at the N6 position within GTAC motifs. Following DNA isolation, DpnI selectively cleaves DNA at GmATC positions. Cleaved DNA is ligated to primers for PCR amplification. After sufficient amplification, DNA is sent for sequencing. (C) ChEC-sequencing. MNase-candidate TF fusion binds to DNA. Cells are permeabilized, and Ca2+ is added, causing MNase to cleave nearby DNA. Cleaved DNA is isolated and ligated to primers for PCR amplification. After sufficient amplification, DNA is sent for sequencing. (D) CUT&Tag. Cells are isolated and permeabilized, with primary antibody added to bind to target protein and secondary antibody added to increase yields. A Protein-A–Tn5 fusion protein is added, binding to the secondary antibody. Mg2+ is added to activate the Tn5. Tn5 cleaves adjacent DNA and ligates sequencing adapters to DNA. SDS-PAGE is added to the reaction to release DNA fragments, ready for sequencing.











