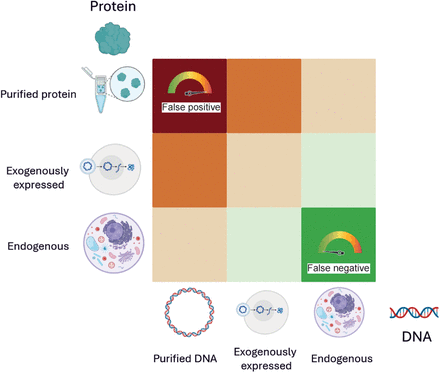
Matrix of biological relevance in DNA–protein interaction studies. Experimental approaches to study DNA–protein interactions vary in their use of cis-elements (DNA) and trans-elements (proteins). The matrix illustrates combinations of DNA substrates (in vitro naked DNA, exogenously introduced DNA, and endogenous chromatin) and protein sources (purified, plasmid-expressed, or endogenously produced) arranged by increasing physiological relevance along the diagonal from bottom right to top left. Cells are color-coded to reflect biological relevance, ranging from low (red) to high (green). Systems using purified proteins and naked DNA are prone to false positives, whereas fully endogenous systems, although biologically informative, carry a higher risk of false negatives owing to contextual dependencies such as chromatin state, cell cycle stage, and transcriptional noise. It is critical to carefully select experimental configurations to balance control, sensitivity, and physiological accuracy when investigating transcription factor–DNA interactions, although of course no method is free of risk; for example, EMSA can also generate false negatives (e.g., false negative for binding when a cofactor would be required in the cell environment).











