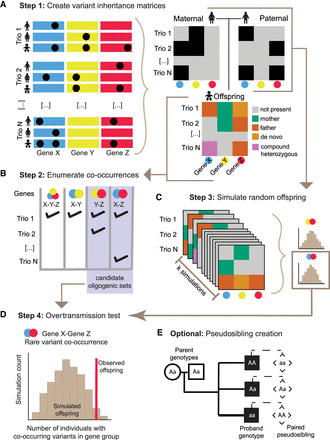
The GCOD approach identifies sets of genes that interact in oligogenic disease. (A) Step 1. Users submit a list of variants of interest or define criteria for some level of “rare” or “predicted damaging.” These variants are summarized in two binary gene-by-trio matrices that denote whether at least one damaging allele was present in each of the maternal and paternal genomes, as well as an offspring matrix that encodes damaging variant presence and provenance. The variant(s) in each gene can be only maternally derived, only paternally derived, compound heterozygous, or de novo. (B) Step 2. From the offspring matrix, GCOD enumerates the candidate sets of genes for which variants occur in multiple probands, including only combinations not transmitted from a single parent. (C) Step 3. An offspring is created from the genotypes of each pair of parents over a minimum of 10,000 simulated cohorts, yielding a distribution of simulated co-occurrence counts for each candidate set. (D) Step 4. For a given candidate set, the simulated distribution is compared to the observed number in cohort probands to compute a P-value. (E) Optionally, GCOD derives pseudosibling genotypes from the parental alleles not transmitted to the proband for comparative analysis.











