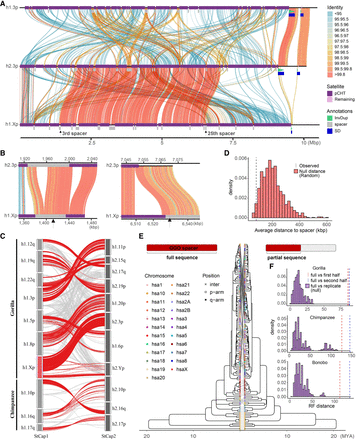
Evidence of nonallelic sequence exchange at SD spacers. (A) An alignment view (SVbyEye) (Porubsky et al. 2024) depicting a candidate ectopic exchange between subterminal satellite chromosomes hsa3 and hsaX p-arms of gorilla. The percentage identity of alignment (scaled from blue <95% to red >99.8%) with annotation tracks for higher-order pCht blocks, satellites, and SDs is shown on the right. (B) Enlarged view of the two exchange breakpoints (black arrows) mapping within the third and 25th SD spacers. (C) Overview of potential ectopic exchange events between the subterminal caps (left-StCap1 vs. right-StCap2 tracks) identified in African great ape genomes (eight and three events in gorilla and chimpanzee, respectively, including hsa3p vs. hsaXp in red). Assembly quality at the breakpoints was validated by a read-depth analysis in Supplemental Figures S20–S26. (D) Simulation test of exchange event breakpoints, suggesting that the exchange breaks (>99.8% identity alignment blocks; dotted line) are more likely to be located within or close to subterminal SD spacers compared with a random distribution (red histogram). (E,F) Comparison of maximum likelihood phylogeny to test recombination of SD spacer. A phylogenetic tree of the complete spacer sequence (left) is compared with that of a subset of the sequence (right). The tip nodes of the left tree are linked to the corresponding nodes of the right tree to visualize phylogenetic shifts in the topology. Robinson–Foulds (RF) distance is used to measure the extent of phylogenetic shift (first or last half sequences, in red and blue dotted lines, compared to a null distribution generated by computing distances of consensus tree vs. bootstrap replicate trees).











