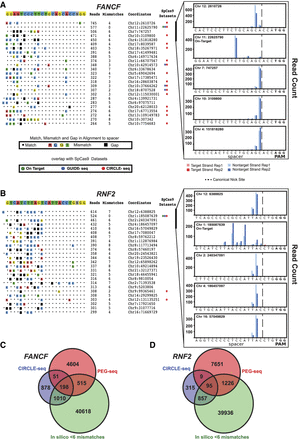
PEG-seq can detect in vitro off-target nicking by the prime editor. (A,B, left) Sorted list of top 25 highest signal peaks in PEG-seq data for FANCF (A) or RNF2 (B) sgRNA spacer sequences. For each site, the alignment to the spacer sequence is indicated with any mismatches (colored boxes) or gaps (black boxes), average read number, genomic location, and overlap with SpCas9 off-target data sets for CIRCLE-seq or GUIDE-seq shown for each potential off-target site. (Right) Plots of the top five sites for each spacer are shown for two replicates (indicated by shading) displaying the accumulation of DNA breaks across the spacer sequence with strand and base pair resolution. It is notable that the precise position of the reads relative to the expected canonical nick site (gapped line) varies across sites and is notably left-shifted at the on-target site. (C,D) Venn diagrams of genome-wide sites identified by PEG-seq, CIRCLE-seq, and in silico analysis (sites six or fewer mismatches or gaps in the human genome) for the FANCF (C) and RNF2 (D) spacers.











