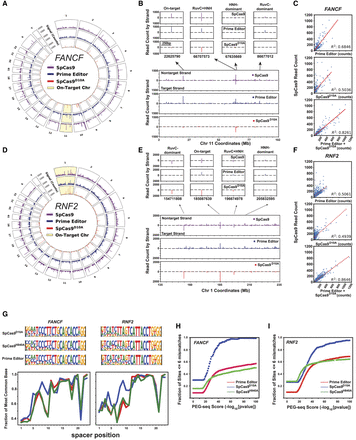
PEG-seq protocol detects off-target SSBs generated by SpCas9, prime editor, and SpCas9D10A genome-wide. (A,D) Circle plot of genome-wide PEG-seq signal for each indicated enzyme for the FANCF (A) or RNF2 (D) guide RNA. (B,E) Zoom-in on 100 Mb of Chromosome 11 showing the on-target site for FANCF guide (B) or 100 Mb on Chromosome 1 for the RNF2 guide (E) and the off-target signal that is apparent in enzymes with only the RuvC (prime editor) or only HNH activity (SpCas9D10A) or both (SpCas9). (C,F) Analysis of SpCas9 signal plotted against either prime editor, SpCas9D10A counts, or the sum of the PE and SpCas9D10A counts for FANCF (C) or RNF2 (F) guides. As expected the SpCas9 signal is strongly predicted by the sum of the signals of the two individual enzymatic activities. (G, top) Position-weight matrices for all sites identified with PEG-seq using the FANCF or RNF2 sgRNA and prime editor, SpCas9D10A, or SpCas9H840A proteins. (Bottom) Line plots of the most frequent bases at each position in the spacer and PAM for the top 100 sites as identified by PEG-seq for FANCF and RNF2. (H,I) Plots of fraction of sites identified by PEG-seq at each score threshold that are within six mismatches or gaps of the indicated spacer sequence.











