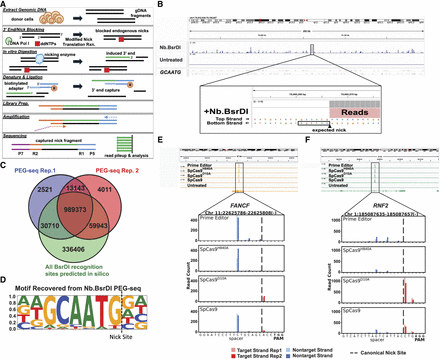
Development and validation of PEG-seq as a genome-wide nick detection method. (A) Diagram of PEG-seq protocol for capture of single-strand breaks (SSBs) in DNA induced by an enzyme. (B) PEG-seq detection of nicks from Nb.BsrDI-treated DNA visualized using IGV (Robinson et al. 2011) with a ∼280 kb region view and zoom-in (52 bp) to one peak showing a pileup of reads next to the Nb.BsrDI enzyme recognition site. (C) Overlap (dark brown) between Nb.BsrD1 PEG-seq replicates and sites identified by in silico analysis across the genome. (D) The motif identified from PEG-seq sites is identical to that of the Nb.BsrD1 recognition site (GCAATG). (E,F) Identification of peaks in PEG-seq data at the on-target site for the prime editor, SpCas9H840A, SpCas9D10A, and SpCas9 complexed with sgRNA targeting FANCF (E) or RNF2 (F) sequences. (Top) Genomic region around the target site in IGV indicating one significant peak in all treated conditions. (Bottom) Zoomed-in view of signal around the spacer sequence of the on-target site; reads from two replicates are shown with base pair resolution (indicated by dark and light shading). As expected, reads from the SpCas9D10A data set are on the target strand, and prime editor or SpCas9H840A shows accumulation of reads on the nontarget strand; SpCas9 shows reads on both strands as it maintains both enzymatic activities. The expected site of canonical nicking is indicated with a dashed line and aligns with the target strand (SpCas9D10A) signal, whereas the nontarget strand signal (prime editor, SpCas9H840A, SpCas9) is offset by 4–6 nucleotides.











