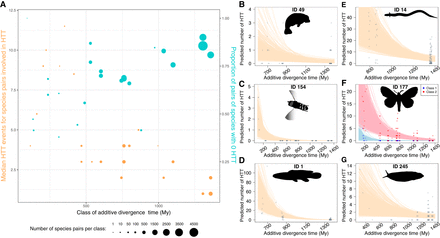
Phylogenetic relatedness favors horizontal transfer of transposable elements. (A) Correlation between divergence time and (i) the median number of HTT events per pair of species, excluding pairs not involved in HTT (orange circles, left y-axis, r = 0.69), and (ii) the proportion of pairs of species in which no HTT was inferred (turquoise circles, right y-axis, r = –0.72). Both (i) and (ii) were computed per class of divergence time, which are 100 My a.d.t. wide and overlap by 50 My a.d.t., using one species per species unit (Fig. 2B). Both correlation coefficients are weighted using the natural logarithm of the number of species pairs composing each class, to account for more precise estimates when many species pairs compose the median or proportion (here, the higher the a.d.t., the more species pairs are in the class of additive divergence time). Circles are horizontally located at the median divergence times within classes. (B–G) Estimates from a negative binomial regression model of the number of HTT events in a focal species as a function of the divergence time from “partner” species. IDs of the focal species are indicated in each panel; they correspond to tips in Figure 2B and their corresponding species names can be found in Supplemental Dataset S1. The model estimates are provided with the assumption of shared habitat. In F, Class 1 and Class 2 TEs are modeled separately. Semitransparent lines correspond to the expected number of HTT estimated from each of 36,000 Markov chain Monte Carlo draws. Points represent observed data and are slightly randomly shifted on the x-axis to minimize overlapping.











